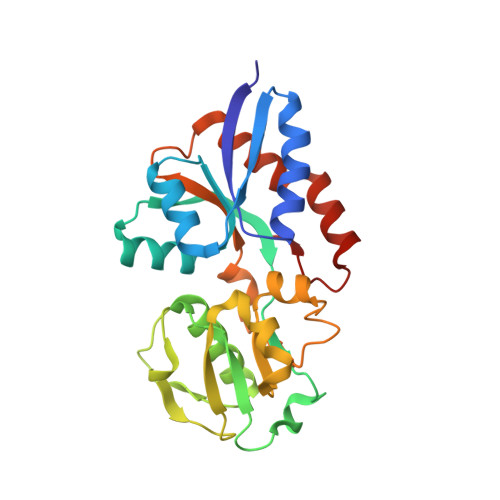
Structural evidence that the 32-kilodalton lipoprotein (Tp32) of Treponema pallidum is an L-methionine-binding protein
Deka, R.K., Neil, L., Hagman, K.E., Machius, M., Tomchick, D.R., Brautigam, C.A., Norgard, M.V.(2004) J Biological Chem 279: 55644-55650
- PubMed: 15489229 Search on PubMed
- DOI: https://doi.org/10.1074/jbc.M409263200
- Primary Citation Related Structures:
1XS5 - PubMed Abstract:
A structure-to-function approach was undertaken to gain insights into the potential function of the 32-kDa membrane lipoprotein (Tp32) of Treponema pallidum, the syphilis bacterium. The crystal structure of rTp32 (determined at a resolution of 1.85 A) shows that the organization of rTp32 is similar to other periplasmic ligand-binding proteins (PLBPs), in that it consists of two alpha/beta domains, linked by two crossovers, with a binding pocket between them. In the pocket, a molecule of L-methionine was detected in the electron density map. Residues from both domains interact with the ligand. One of the crossover regions is comprised of a 3(10)-helix, a feature not typical in other ligand-binding proteins. Sequence comparison shows strong similarity to other hypothetical methionine-binding proteins. Together, the data support the notion that rTp32 is a component of a periplasmic methionine uptake transporter system in T. pallidum.
- Department of Microbiology, University of Texas Southwestern Medical Center, 6000 Harry Hines Boulevard, Dallas, TX 75390, USA.
Organizational Affiliation: