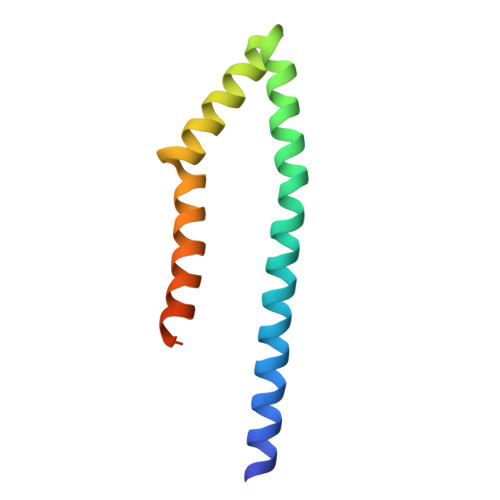
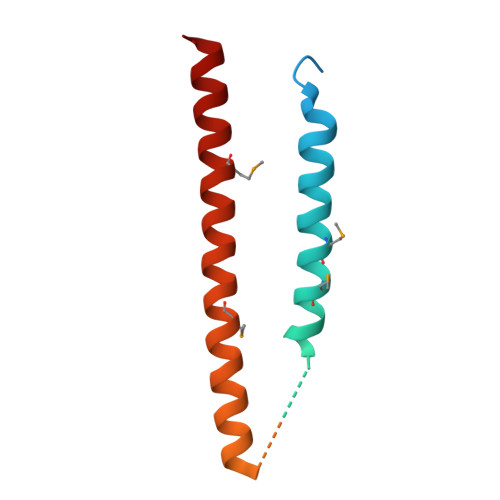
Structural characterization of a type III secretion system filament protein in complex with its chaperone.
Yip, C.K., Finlay, B.B., Strynadka, N.C.J.(2005) Nat Struct Mol Biol 12: 75-81
- PubMed: 15619638 Search on PubMed
- DOI: https://doi.org/10.1038/nsmb879
- Primary Citation Related Structures:
1XOU - PubMed Abstract:
The type III secretion system (TTSS) mediates the specific translocation of bacterial proteins into the cytoplasm of eukaryotic cells, a process essential for the virulence of many Gram-negative pathogens. The enteropathogenic Escherichia coli TTSS protein EspA forms a hollow extracellular filament believed to be a molecular conduit for type III protein translocation. Structural analysis of EspA has been hampered by its polymeric nature. We show that EspA alone is sufficient to form filamentous structures in the absence of other pathogenicity island-encoded proteins. CesA is the recently proposed chaperone of EspA, and we demonstrate that CesA traps EspA in a monomeric state and inhibits its polymerization. Crystallographic analysis of the heterodimeric CesA-EspA complex at a resolution of 2.8 A reveals that EspA contains two long a-helices, which are involved in extensive coiled-coil interactions with CesA.
- Department of Biochemistry and Molecular Biology, University of British Columbia, 2146 Health Sciences Mall, Vancouver, British Columbia, Canada V6T 1Z3.
Organizational Affiliation: