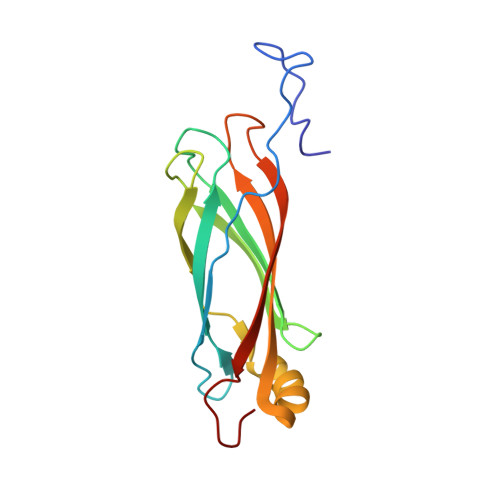
Solution structure of a late embryogenesis abundant protein (LEA14) from Arabidopsis thaliana, a cellular stress-related protein
Singh, S., Cornilescu, C.C., Tyler, R.C., Cornilescu, G., Tonelli, M., Lee, M.S., Markley, J.L.(2005) Protein Sci 14: 2601-2609
- PubMed: 16155204 Search on PubMedSearch on PubMed Central
- DOI: https://doi.org/10.1110/ps.051579205
- Primary Citation Related Structures:
1XO8 - PubMed Abstract:
We report the three-dimensional structure of a late embryogenesis abundant (LEA) protein from Arabidopsis thaliana gene At1g01470.1. This protein is a member of Pfam cluster PF03168, and has been classified as a LEA14 protein. LEA proteins are expressed under conditions of cellular stress, such as desiccation, cold, osmotic stress, and heat. The structure, which was determined by NMR spectroscopy, revealed that the At1g01470.1 protein has an alphabeta-fold consisting of one alpha-helix and seven beta-strands that form two antiparallel beta-sheets. The closest structural homologs were discovered to be fibronectin Type III domains, which have <7% sequence identity. Because fibronectins from animal cells have been shown to be involved in cell adhesion, cell motility, wound healing, and maintenance of cell shape, it is interesting to note that in plants wounding or stress results in the overexpression of a protein with fibronectin Type III structural features.
- Center for Eukaryotic Structural Genomics, University of Wisconsin-Madison, 433 Babcock Drive, Madison, WI 53706, USA.
Organizational Affiliation: