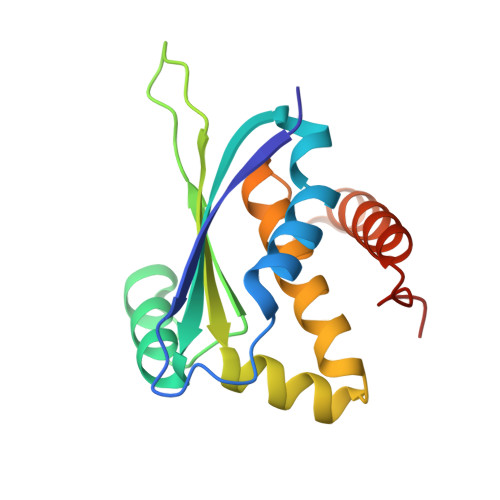
The ybeY protein from Escherichia coli is a metalloprotein.
Zhan, C., Fedorov, E.V., Shi, W., Ramagopal, U.A., Thirumuruhan, R., Manjasetty, B.A., Almo, S.C., Fiser, A., Chance, M.R., Fedorov, A.A.(2005) Acta Crystallogr Sect F Struct Biol Cryst Commun 61: 959-963
- PubMed: 16511207 Search on PubMedSearch on PubMed Central
- DOI: https://doi.org/10.1107/S1744309105031131
- Primary Citation Related Structures:
1XM5 - PubMed Abstract:
The three-dimensional crystallographic structure of the ybeY protein from Escherichia coli (SwissProt entry P77385) is reported at 2.7 A resolution. YbeY is a hypothetical protein that belongs to the UPF0054 family. The structure reveals that the protein binds a metal ion in a tetrahedral geometry. Three coordination sites are provided by histidine residues, while the fourth might be a water molecule that is not seen in the diffraction map because of its relatively low resolution. X-ray fluorescence analysis of the purified protein suggests that the metal is a nickel ion. The structure of ybeY and its sequence similarity to a number of predicted metal-dependent hydrolases provides a functional assignment for this protein family. The figures and tables of this paper were prepared using semi-automated tools, termed the Autopublish server, developed by the New York Structural GenomiX Research Consortium, with the goal of facilitating the rapid publication of crystallographic structures that emanate from worldwide Structural Genomics efforts, including the NIH-funded Protein Structure Initiative.
- New York Structural Genomics Research Consortium (NYSGXRC), Albert Einstein College of Medicine, Bronx, New York 10461, USA.
Organizational Affiliation: