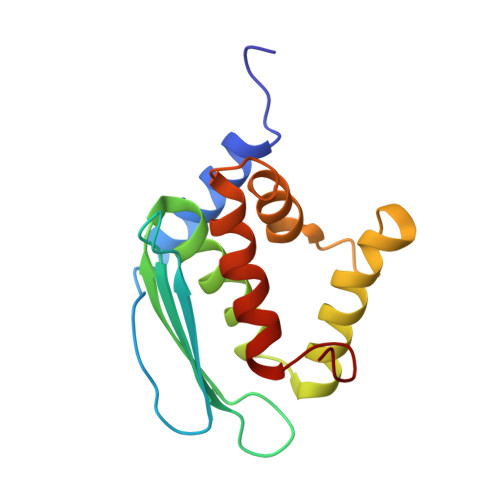
Solution structure of Iron-Sulfur cluster assembly protein IscU from Bacillus subtilis, with Zinc bound at the active site. Northeast Structural Genomics Consortium Target SR17.
Kornhaber, G.J., Swapna, G.V.T., Ramelot, T.A., Cort, J.R., Kennedy, M.A., Montelione, G.T.To be published.