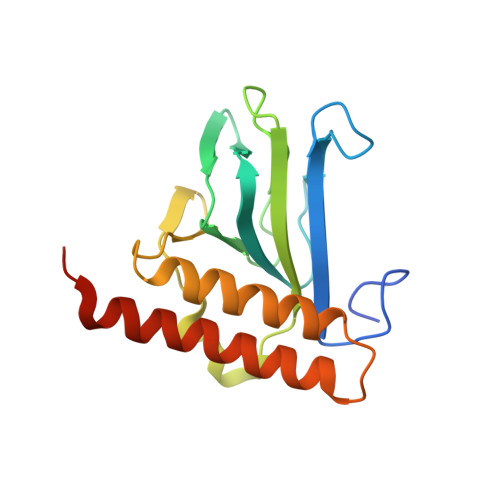
Structure and properties of a truely apo form of AraC dimerization domain.
Weldon, J.E., Rodgers, M.E., Larkin, C., Schleif, R.F.(2007) Proteins 66: 646-654
- PubMed: 17173282 Search on PubMed
- DOI: https://doi.org/10.1002/prot.21267
- Primary Citation Related Structures:
1XJA - PubMed Abstract:
The arabinose-binding pockets of wild type AraC dimerization domains crystallized in the absence of arabinose are occupied with the side chains of Y31 from neighboring domains. This interaction leads to aggregation at high solution concentrations and prevents determination of the structure of truely apo AraC. In this work we found that the aggregation does not significantly occur at physiological concentrations of AraC. We also found that the Y31V mutation eliminates the self-association, but does not affect regulation properties of the protein. At the same time, the mutation allows crystallization of the dimerization domain of the protein with only solvent in the arabinose-binding pocket. Using a distance difference method suitable for detecting and displaying even minor structural variation among large groups of similar structures, we find that there is no significant structural change in the core of monomers of the AraC dimerization domain resulting from arabinose, fucose, or tyrosine occupancy of the ligand-binding pocket. A slight change is observed in the relative orientation of monomers in the dimeric form of the domain upon the binding of arabinose but its significance cannot yet be assessed.
- Department of Biology, Johns Hopkins University, Baltimore, Maryland 21218, USA.
Organizational Affiliation: