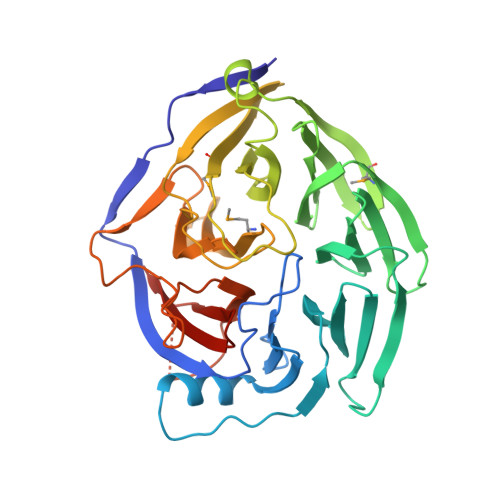
The N-Terminal Domain of Nup159 Forms a beta-Propeller that Functions in mRNA Export by Tethering the Helicase Dbp5 to the Nuclear Pore
Weirich, C.S., Erzberger, J.P., Berger, J.M., Weis, K.(2004) Mol Cell 16: 749-760
- PubMed: 15574330 Search on PubMed
- DOI: https://doi.org/10.1016/j.molcel.2004.10.032
- Primary Citation Related Structures:
1XIP - PubMed Abstract:
Nuclear export of mRNA in eukaryotic cells is mediated by soluble transport factors and components of the nuclear pore complex (NPC). The cytoplasmically oriented nuclear pore protein Nup159 plays a critical role in mRNA export through its conserved N-terminal domain (NTD). Here, we report the crystal structure of the Nup159 NTD, refined to 2.5 A. The structure reveals an unusually asymmetric seven-bladed beta-propeller that is structurally conserved throughout eukarya. Using structure-based conservation analysis, we have targeted specific surface residues for mutagenesis. Residue substitutions in a conserved loop of the NTD abolish in vitro binding to Dbp5, a DEAD box helicase required for mRNA export. In vivo, these mutations cause Dbp5 mislocalization and block mRNA export. These findings suggest that the Nup159 NTD functions in mRNA export as a binding platform, tethering shuttling Dbp5 molecules at the nuclear periphery and locally concentrating this mRNA remodeling factor at the cytoplasmic face of the NPC.
- Division of Cell and Developmental Biology, Department of Molecular and Cell Biology, University of California, Berkeley, Berkeley, California 94720, USA.
Organizational Affiliation: