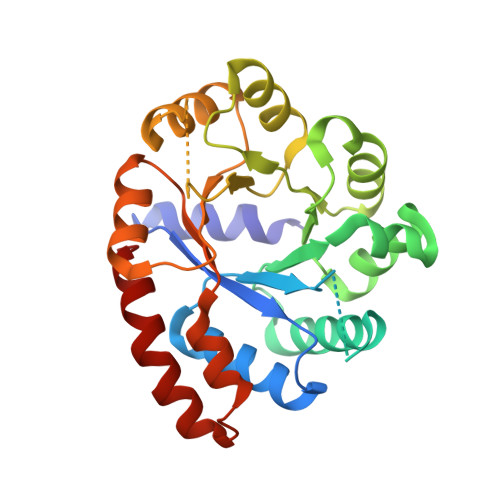
Structures of wild-type and P28L/Y173F tryptophan synthase alpha-subunits from Escherichia coli
Jeong, M.S., Jeong, J.K., Lim, W.K., Jang, S.B.(2004) Biochem Biophys Res Commun 323: 1257-1264
- PubMed: 15451433
- DOI: https://doi.org/10.1016/j.bbrc.2004.08.222
- Primary Citation Related Structures:
1XC4, 1XCF - PubMed Abstract:
The alpha-subunit of tryptophan synthase (alphaTS) catalyzes the cleavage of indole-3-glycerol phosphate to glyceraldehyde-3-phosphate and indole, which is used to yield the amino acid tryptophan in tryptophan biosynthesis. Here, we report the first crystal structures of wild-type and double-mutant P28L/Y173F alpha-subunit of tryptophan synthase from Escherichia coli at 2.8 and 1.8A resolution, respectively. The structure of wild-type alphaTS from E. coli was similar to that of the alpha(2)beta(2) complex structure from Salmonella typhimurium. As compared with both structures, the conformational changes are mostly in the interface of alpha- and beta-subunits, and the substrate binding region. Two sulfate ions and two glycerol molecules per asymmetric unit bind with the residues in the active sites of the wild-type structure. Contrarily, double-mutant P28L/Y173F structure is highly closed at the window for the substrate binding by the conformational changes. The P28L substitution induces the exposure of hydrophobic amino acids and decreases the secondary structure that causes the aggregation. The Y173F suppresses to transfer a signal from the alpha-subunit core to the alpha-subunit surface involved in interactions with the beta-subunit and increases structural stability.
- Korea Nanobiotechnology Center, Pusan National University, Jangjeon-dong, Geumjeong-gu, Busan 609-735, Republic of Korea.
Organizational Affiliation: