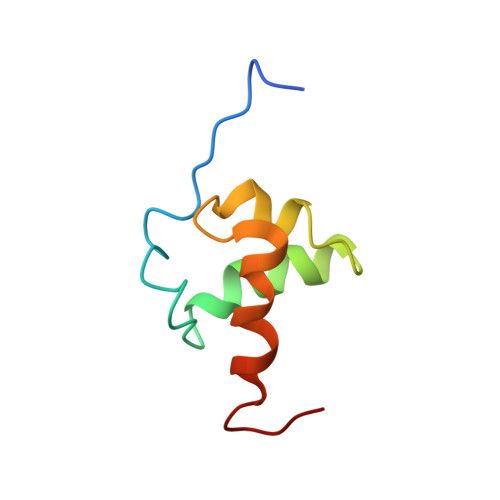
Structural insights into the interaction and activation of histone deacetylase 3 by nuclear receptor corepressors
Codina, A., Love, J.D., Li, Y., Lazar, M.A., Neuhaus, D., Schwabe, J.W.R.(2005) Proc Natl Acad Sci U S A 102: 6009-6014
- PubMed: 15837933 Search on PubMedSearch on PubMed Central
- DOI: https://doi.org/10.1073/pnas.0500299102
- Primary Citation Related Structures:
1XC5 - PubMed Abstract:
SMRT (silencing mediator of retinoid acid and thyroid hormone receptor) and NCoR (nuclear receptor corepressor) are transcriptional corepressors that play an essential role in the regulation of development and metabolism. This role is achieved, in part, through the recruitment of a key histone deacetylase (HDAC3), which is itself indispensable for cell viability. The assembly of HDAC3 with the deacetylase activation domain (DAD) of SMRT and NCoR is required for activation of the otherwise inert deacetylase. The DAD comprises an N-terminal DAD-specific motif and a C-terminal SANT (SWI3/ADA2/NCoR/TFIIIB)-like domain. We report here the solution structure of the DAD from SMRT, which reveals a four-helical structure. The DAD differs from the SANT (and MYB) domains in that (i) it has an additional N-terminal helix and (ii) there is a notable hydrophobic groove on the surface of the domain. Structure-guided mutagenesis, combined with interaction assays, showed that residues in the vicinity of the hydrophobic groove are required for interaction with (and hence activation of) HDAC3. Importantly, one surface-exposed lysine is required for activation of HDAC3, but not for interaction. This lysine may play a uniquely important role in the mechanism of activating HDAC3.
- Medical Research Council, Laboratory of Molecular Biology, Hills Road, Cambridge CB2 2QH, United Kingdom.
Organizational Affiliation: