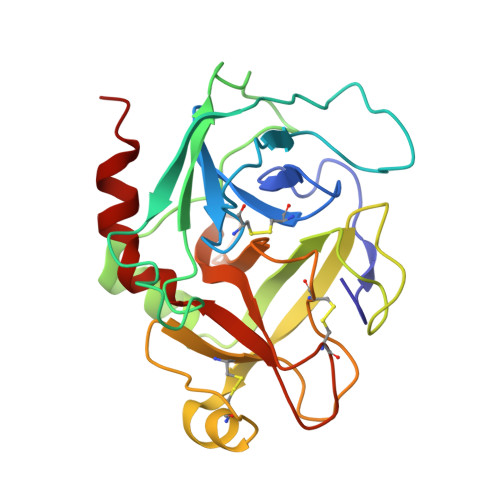
SAR and factor IXa crystal structure of a dual inhibitor of factors IXa and Xa
Smallheer, J.M., Alexander, R.S., Wang, J., Wang, S., Nakajima, S., Rossi, K.A., Smallwood, A., Barbera, F., Burdick, D., Luettgen, J.M., Knabb, R.M., Wexler, R.R., Jadhav, P.K.(2004) Bioorg Med Chem Lett 14: 5263-5267
- PubMed: 15454208 Search on PubMed
- DOI: https://doi.org/10.1016/j.bmcl.2004.08.034
- Primary Citation Related Structures:
1X7A - PubMed Abstract:
Modifications to the P4 moiety and pyrazole C3 substituent of factor Xa inhibitor SN-429 provided several new compounds, which are 5-10nM inhibitors of factor IXa. An X-ray crystal structure of one example complexed to factor IXa shows that these compounds adopt a similar binding mode to that previously observed with pyrazole inhibitors in the factor Xa active site both with regard to how the inhibitor binds and the position of Tyr99.
- Bristol-Myers Squibb Company, PO Box 5400, Princeton, NJ 08543-5400, USA. joanne.smallheer@bms.com
Organizational Affiliation: