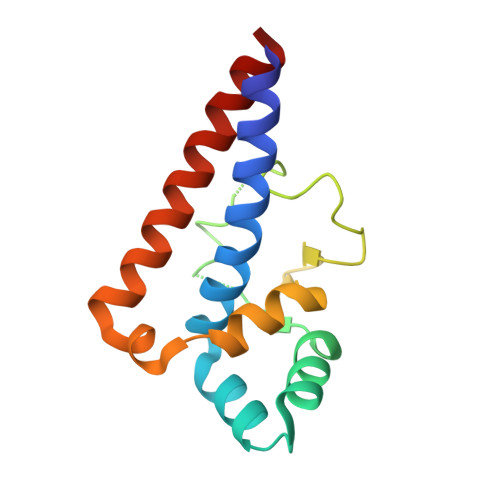
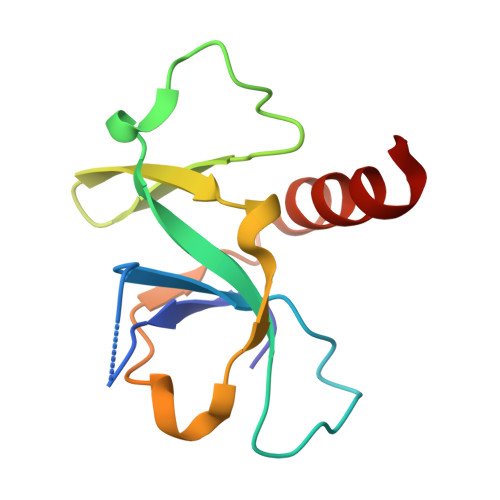
Molecular Basis of Gyrase Poisoning by the Addiction Toxin CcdB
Dao-Thi, M.H., Van Melderen, L., De Genst, E., Afif, H., Buts, L., Wyns, L., Loris, R.(2005) J Mol Biology 348: 1091-1102
- PubMed: 15854646 Search on PubMed
- DOI: https://doi.org/10.1016/j.jmb.2005.03.049
- Primary Citation Related Structures:
1X75 - PubMed Abstract:
Gyrase is an ubiquitous bacterial enzyme that is responsible for disentangling DNA during DNA replication and transcription. It is the target of the toxin CcdB, a paradigm for plasmid addiction systems and related bacterial toxin-antitoxin systems. The crystal structure of CcdB and the dimerization domain of the A subunit of gyrase (GyrA14) dictates an open conformation for the catalytic domain of gyrase when CcdB is bound. The action of CcdB is one of a wedge that stabilizes a dead-end covalent gyrase:DNA adduct. Although CcdB and GyrA14 form a globally symmetric complex where the two 2-fold axes of both dimers align, the complex is asymmetric in its details. At the centre of the interaction site, the Trp99 pair of CcdB stacks with the Arg462 pair of GyrA14, explaining why the Arg462Cys mutation in the A subunit of gyrase confers resistance to CcdB. Overexpression of GyrA14 protects Escherichia coli cells against CcdB, mimicking the action of the antidote CcdA.
- Laboratorium voor Ultrastructuur, Vrije Universiteit Brussel and Department of Molecular and Cellular Interactions, Vlaams Instituut voor Biotechnologie, Building E, Pleinlaan 2, B-1050 Brussels, Belgium.
Organizational Affiliation: