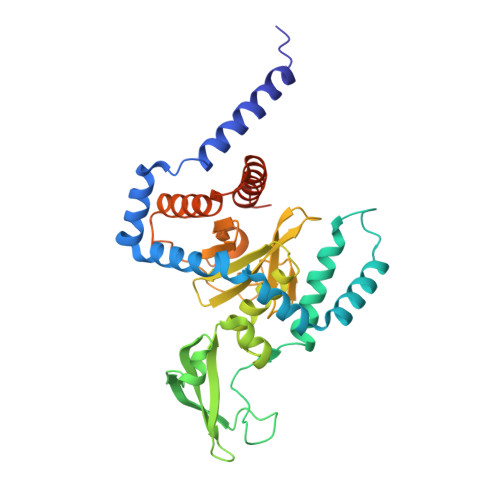
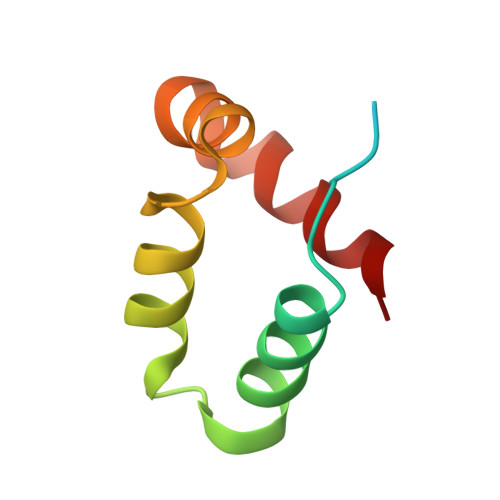
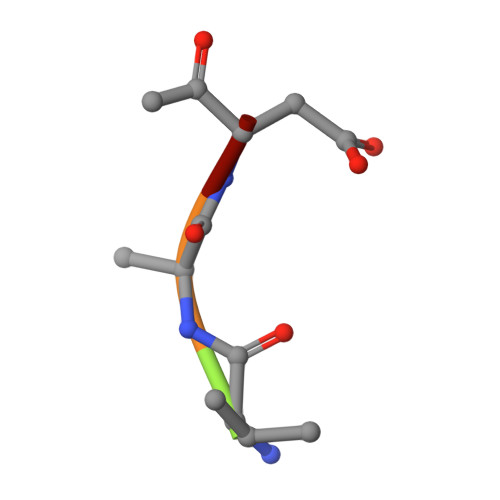
Structure of a peptide:N-glycanase-Rad23 complex: insight into the deglycosylation for denatured glycoproteins.
Lee, J.H., Choi, J.M., Lee, C., Yi, K.J., Cho, Y.(2005) Proc Natl Acad Sci U S A 102: 9144-9149
- PubMed: 15964983 Search on PubMedSearch on PubMed Central
- DOI: https://doi.org/10.1073/pnas.0502082102
- Primary Citation Related Structures:
1X3W, 1X3Z - PubMed Abstract:
In eukaryotes, misfolded proteins must be distinguished from correctly folded proteins during folding and transport processes by quality control systems. Yeast peptide:N-glycanase (yPNGase) specifically deglycosylates the denatured form of N-linked glycoproteins in the cytoplasm and assists proteasome-mediated glycoprotein degradation by forming a complex with 26S proteasome through DNA repair protein, yRad23. Here, we describe the crystal structures of a yPNGase and XPC-binding domain of yRad23 (yRad23XBD, residues 238-309) complex and of a yPNGase-yRad23XBD complex bound to a caspase inhibitor, Z-VAD-fmk. yPNGase is formed with three domains, a core domain containing a Cys-His-Asp triad, a Zn-binding domain, and a Rad23-binding domain. Both N- and C-terminal helices of yPNGase interact with yRad23 through extensive hydrophobic interactions. The active site of yPNGase is located in a deep cleft that is formed with residues conserved in all PNGase members, and three sugar molecules are bound to this cleft. Complex structures in conjunction with mutational analyses revealed that the walls of the cleft block access to the active site of yPNGase by native glycoprotein, whereas the cleft is sufficiently wide to accommodate denatured glycoprotein, thus explaining the specificity of PNGase for denatured substrates.
- National Creative Research Center for Structural Biology and Department of Life Science, Pohang University of Science and Technology, Hyo-ja dong, San 31, Pohang, KyungBook, South Korea.
Organizational Affiliation: