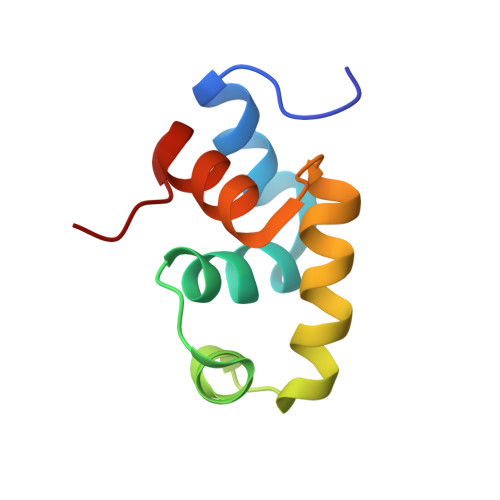
Solution structure of the C-terminal transcriptional activator domain of FixJ from Sinorhizobium meliloti and its recognition of the fixK promoter
Kurashima-Ito, K., Kasai, Y., Hosono, K., Tamura, K., Oue, S., Isogai, M., Ito, Y., Nakamura, H., Shiro, Y.(2005) Biochemistry 44: 14835-14844
- PubMed: 16274231
- DOI: https://doi.org/10.1021/bi0509043
- Primary Citation of Related Structures:
1X3U - PubMed Abstract:
FixJ is a response regulator of the two-component signal transduction pathway involved in the transcriptional activation of nitrogen fixation genes of Sinorhizobium meliloti. Upon phosphorylation, FixJ transcriptionally activates the fixK and nifA promoters. We identified a FixJ recognition sequence of 16 bp in the high affinity binding site of the fixK promoter by means of a gel shift assay. In addition, the solution structure of the truncated C-terminal DNA binding domain of FixJ (FixJC) was solved by NMR spectroscopy. FixJC contains five alpha-helices that encode a typical helix-turn-helix motif as a potential DNA binding core with the highest structural similarity toward the C-terminal DNA binding domain of NarL. The addition of the DNA fragment containing the recognition sequence of the high affinity FixJ binding site resulted in intermediate to slow exchange interactions on the NMR time scale in the spectrum of FixJC, while the exchange was rapid in the case of control DNA. These spectral data suggest that more than one molecule of FixJC binds to the recognition sequence, although FixJC alone is present in monomeric form in solution. This result is consistent with a scenario in which a transcriptionally active species of FixJ is a homodimer of the phosphorylated form.
- Yokohama City University, Suehiro, Tsurumi, Yokohama, Kanagawa 230-0045, Japan.
Organizational Affiliation: