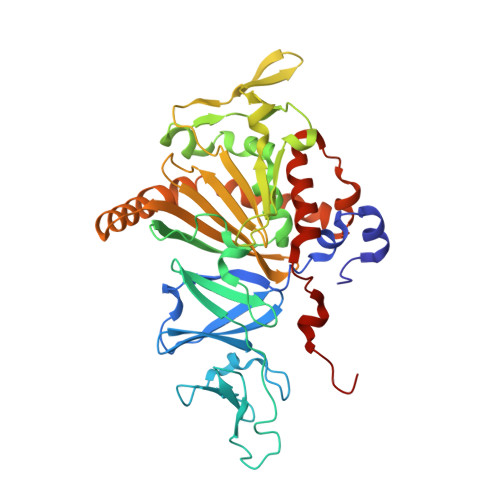
Structure of the terminal oxygenase component of angular dioxygenase, carbazole 1,9a-dioxygenase
Nojiri, H., Ashikawa, Y., Noguchi, H., Nam, J.-W., Urata, M., Fujimoto, Z., Uchimura, H., Terada, T., Nakamura, S., Shimizu, K., Yoshida, T., Habe, H., Omori, T.(2005) J Mol Biology 351: 355-370
- PubMed: 16005887
- DOI: https://doi.org/10.1016/j.jmb.2005.05.059
- Primary Citation of Related Structures:
1WW9 - PubMed Abstract:
Carbazole 1,9a-dioxygenase (CARDO) catalyzes the dihydroxylation of carbazole by angular position (C9a) carbon bonding to the imino nitrogen and its adjacent C1 carbon. This reaction is an initial degradation reaction of the carbazole degradation pathway by various bacterial strains. Only a limited number of Rieske non-heme iron oxygenase systems (ROSs) can catalyze this novel reaction, termed angular dioxygenation. Angular dioxygenation is also involved in the degradation pathways of carbazole-related compounds, dioxin, and CARDO can catalyze the angular dioxygenation for dioxin. CARDO consists of a terminal oxygenase component (CARDO-O), and the electron transport components, ferredoxin (CARDO-F) and ferredoxin reductase (CARDO-R). CARDO-O has a homotrimeric structure, and governs the substrate specificity of CARDO. Here, we have determined the crystal structure of CARDO-O of Janthinobacterium sp. strain J3 at a resolution of 1.95A. The alpha3 trimeric overall structure of the CARDO-O molecule roughly corresponds to the alpha3 partial structures of other terminal oxygenase components of ROSs that have the alpha3beta3 configuration. The CARDO-O structure is a first example of the terminal oxygenase components of ROSs that have the alpha3 configuration, and revealed the presence of the specific loops that interact with a neighboring subunit, which is proposed to be indispensable for stable alpha3 interactions without structural beta subunits. The shape of the substrate-binding pocket of CARDO-O is markedly different from those of other oxygenase components involved in naphthalene and biphenyl degradation pathways. Docking simulations suggested that carbazole binds to the substrate-binding pocket in a manner suitable for catalysis of angular dioxygenation.
- Biotechnology Research Center, The University of Tokyo, 1-1-1 Yayoi, Bunkyo-ku, Tokyo 113-8657, Japan. anojiri@mail.ecc.u-tokyo.ac.jp
Organizational Affiliation: