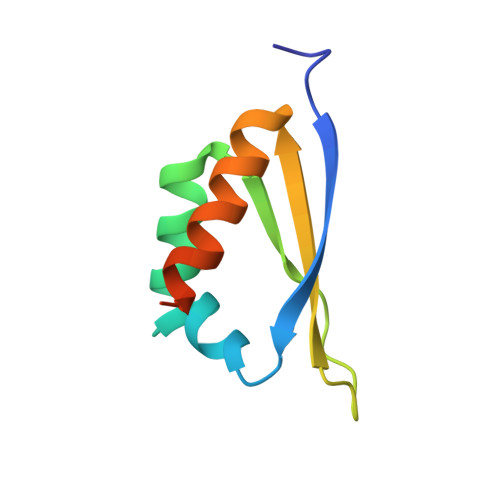
Structure and RNA binding of the third KH domain of poly(C)-binding protein 1.
Sidiqi, M., Wilce, J.A., Vivian, J.P., Porter, C.J., Barker, A., Leedman, P.J., Wilce, M.C.J.(2005) Nucleic Acids Res 33: 1213-1221
- PubMed: 15731341 Search on PubMedSearch on PubMed Central
- DOI: https://doi.org/10.1093/nar/gki265
- Primary Citation Related Structures:
1WVN - PubMed Abstract:
Poly(C)-binding proteins (CPs) are important regulators of mRNA stability and translational regulation. They recognize C-rich RNA through their triple KH (hn RNP K homology) domain structures and are thought to carry out their function though direct protection of mRNA sites as well as through interactions with other RNA-binding proteins. We report the crystallographically derived structure of the third domain of alphaCP1 to 2.1 A resolution. alphaCP1-KH3 assumes a classical type I KH domain fold with a triple-stranded beta-sheet held against a three-helix cluster in a betaalphaalphabetabetaalpha configuration. Its binding affinity to an RNA sequence from the 3'-untranslated region (3'-UTR) of androgen receptor mRNA was determined using surface plasmon resonance, giving a K(d) of 4.37 microM, which is indicative of intermediate binding. A model of alphaCP1-KH3 with poly(C)-RNA was generated by homology to a recently reported RNA-bound KH domain structure and suggests the molecular basis for oligonucleotide binding and poly(C)-RNA specificity.
- School of Biomedical and Chemical Sciences, the UWA Centre for Medical Research, The University of Western Australia WA Australia 6009.
Organizational Affiliation: