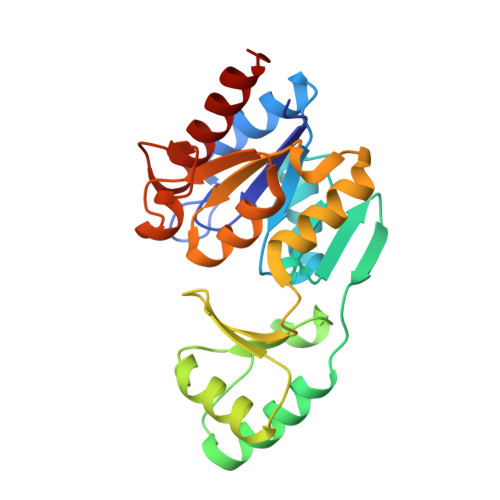
Structure of a haloacid dehalogenase superfamily phosphatase PH1421 from Pyrococcus horikoshii OT3: oligomeric state and thermoadaptation mechanism.
Yamamoto, H., Takio, K., Sugahara, M., Kunishima, N.(2008) Acta Crystallogr D Biol Crystallogr 64: 1068-1077
- PubMed: 18931414 Search on PubMed
- DOI: https://doi.org/10.1107/S0907444908025948
- Primary Citation Related Structures:
1WR8 - PubMed Abstract:
PH1421 from the hyperthermophilic archaeon Pyrococcus horikoshii OT3 is a hypothetical protein belonging to the haloacid dehalogenase (HAD) superfamily. To gain insight into its biological function and thermostabilization mechanism, the crystal structure of PH1421 has been determined at 1.6 A resolution. The crystallographic asymmetric unit contains a homodimer. The monomeric protomer is composed of two distinct domains, a small cap domain and a large core domain, which agrees well with the typical domain organization of HAD subfamily II. Based on structure-based amino-acid sequence alignment and enzymatic analysis, PH1421 is suggested to be a magnesium-dependent phosphatase that is similar to the dimeric HAD phosphatase TA0175 from the mesothermophilic archaeon Thermoplasma acidophilum. Further comparison between the crystal structures of PH1421 and TA0175 revealed a marked structural similarity in the interprotomer dimer association. The common dimer interface with interprotomer twofold symmetry is characterized by a well conserved hydrophobic core consisting of the beta1-alpha1 loop and helices alpha1 and alpha2 of the core domain and additional contacts including the beta7-beta8 loop of the cap domain, which constitutes part of the putative active site of the enzyme. Several factors that potentially contribute to the higher thermal stability of PH1421 were identified: (i) an increase in intraprotomer hydrophobic interactions, (ii) a decrease in denaturation entropy from amino-acid composition and (iii) an increased number of intraprotomer ion pairs. These results suggest that the PH1421 protomer itself has an intrinsically higher thermal stability when compared with the mesothermophilic orthologue TA0175.
- RIKEN SPring-8 Center, Harima Institute, 1-1-1 Kouto, Sayo-cho, Sayo-gun, Hyogo 679-5148, Japan.
Organizational Affiliation: