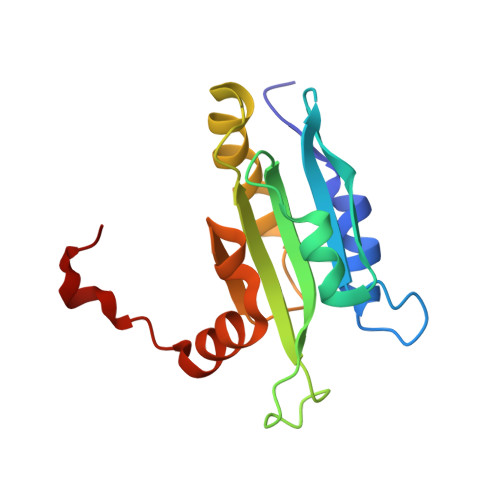
Solution structure of coactosin reveals structural homology to ADF/cofilin family proteins
Hellman, M., Paavilainen, V.O., Naumanen, P., Lappalainen, P., Annila, A., Permi, P.(2004) FEBS Lett 576: 91-96
- PubMed: 15474017 Search on PubMed
- DOI: https://doi.org/10.1016/j.febslet.2004.08.068
- Primary Citation Related Structures:
1WM4 - PubMed Abstract:
Coactosin is a small (MW approximately 15 kDa) evolutionarily conserved actin filament binding protein. It displays remote sequence homology to ADF/cofilin proteins and to the ADF-H domains of twinfilin and Abp1/drebrin. However, biochemical analyses have demonstrated that coactosin has a very different role in actin dynamics from the ones of ADF/cofilin, twinfilin or Abp1/drebrin. To elucidate the molecular mechanism of coactosin/actin interaction, we determined the three-dimensional structure of mouse coactosin by multidimensional NMR spectroscopy. We find that the coactosin structure is homologous to ADF/cofilin and to the ADF-H domains of twinfilin. Furthermore, the regions that have been shown to be important for actin filament interactions in ADF/cofilins are structurally conserved in coactosin suggesting that these two proteins interact with F-actin through a conserved interface. Our analysis also identifies key structural differences between these proteins that may account for the differences in biochemical activities and cellular roles of these proteins.
- Program in Structural Biology and Biophysics, Institute of Biotechnology, University of Helsinki, Finland.
Organizational Affiliation: