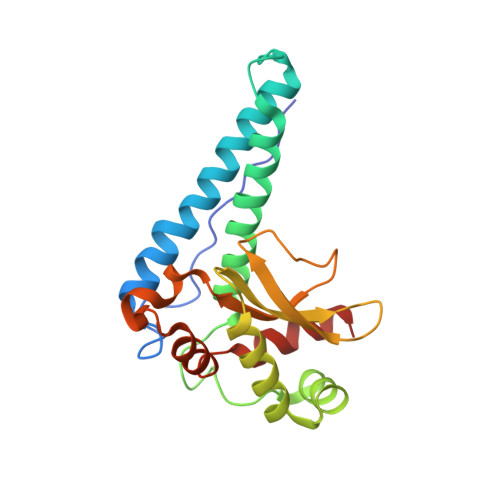
Iron Superoxide Dismutase from the Archaeon Sulfolobus Solfataricus: Analysis of Structure and Thermostability
Ursby, T., Adinolfi, B.S., Al-Karadaghi, S., De Vendittis, E., Bocchini, V.(1999) J Mol Biology 286: 189
- PubMed: 9931259 Search on PubMed
- DOI: https://doi.org/10.1006/jmbi.1998.2471
- Primary Citation Related Structures:
1WB8 - PubMed Abstract:
The crystal structure of superoxide dismutase (SOD) from the hyper thermophile Sulfolobus solfataricus has been determined at 2.3 A resolution by molecular replacement and refined to a crystallographic R-factor of 16.8 % (Rfree 19.8 %). The crystals belong to the space group C2 (a=76.3 A, b=124.3 A, c=60.3 A, beta=128.8 degrees) with two identical monomers in the asymmetric unit. The monomer has a molecular weight of 24 kDa and consists of 210 amino acid residues of which 205 are visible in the electron density map. The overall fold of the monomer of S. solfataricus SOD is similar to that of the other known Fe or Mn-SODs. S. solfataricus SOD forms a very compact tetramer of a type similar to that of SOD from the hyperthermophile Aquifex pyrophilus. Both structures show an elevated number of inter-subunit ion-pairs compared with the mesophilic SOD from Mycobacterium tuberculosis and the thermophilic SOD from Thermus thermophilus. However, in contrast to the A. pyrophilus SOD structure, the number of intra-subunit ion-pairs as well as inter- subunit hydrogen bonds is not higher than in the compared mesophilic and thermophilic SOD structures. The electron density also revealed an unexpected and unusual covalent modification of a conserved tyrosine in the active site. Its involvement in the specific activity of the enzyme is discussed.
- Molecular Biophysics, Center for Chemistry and Chemical Engineering, Lund University, Lund, S-22100, Sweden.
Organizational Affiliation: