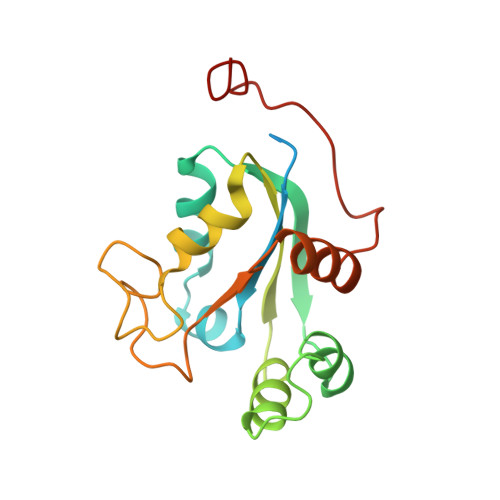
Structure and Mutational Analysis of a Plant Mitochondrial Nucleoside Diphosphate Kinase: Identification of Residues Involved in Serine Phosphorylation and Oligomerisation
Johansson, M., Mackenzie-Hose, A., Andersson, I., Knorpp, C.(2004) Plant Physiol 136: 3034
- PubMed: 15466238 Search on PubMedSearch on PubMed Central
- DOI: https://doi.org/10.1104/pp.104.044040
- Primary Citation Related Structures:
1W7W - PubMed Abstract:
We report the first crystal structure of a plant (Pisum sativum L. cv Oregon sugarpod) mitochondrial nucleoside diphosphate kinase. Similar to other eukaryotic nucleoside diphosphate kinases, the plant enzyme is a hexamer; the six monomers in the asymmetric unit are arranged as trimers of dimers. Different functions of the kinase have been correlated with the oligomeric structure and the phosphorylation of Ser residues. We show that the occurrence of Ser autophosphorylation depends on enzymatic activity. The mutation of the strictly conserved Ser-119 to Ala reduced the Ser phosphorylation to about one-half of that observed in wild type with only a modest change of enzyme activity. We also show that mutating another strictly conserved Ser, Ser-69, to Ala reduces the enzyme activity to 6% and 14% of wild-type using dCDP and dTDP as acceptors, respectively. Changes in the oligomerization pattern of the S69A mutant were observed by cross-linking experiments. A reduction in trimer formation and a change in the dimer interaction could be detected with a concomitant increase of tetramers. We conclude that the S69 mutant is involved in the stabilization of the oligomeric state of this plant nucleoside diphosphate kinase.
- Department of Plant Biology and Forest Genetics, Swedish University of Agricultural Sciences, S-750 07 Uppsala, Sweden.
Organizational Affiliation: