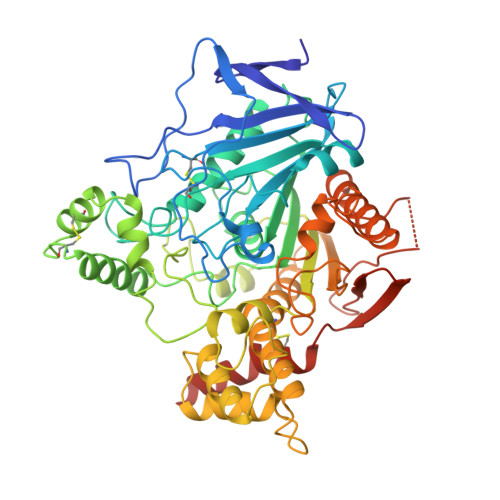
The complex of a bivalent derivative of galanthamine with torpedo acetylcholinesterase displays drastic deformation of the active-site gorge: implications for structure-based drug design.
Greenblatt, H.M., Guillou, C., Guenard, D., Argaman, A., Botti, S., Badet, B., Thal, C., Silman, I., Sussman, J.L.(2004) J Am Chem Soc 126: 15405-15411
- PubMed: 15563167 Search on PubMed
- DOI: https://doi.org/10.1021/ja0466154
- Primary Citation Related Structures:
1W4L, 1W6R, 1W75, 1W76 - PubMed Abstract:
Bifunctional derivatives of the alkaloid galanthamine, designed to interact with both the active site of the enzyme acetylcholinesterase (AChE) and its peripheral cation binding site, have been assayed with Torpedo californica AChE (TcAChE), and the three-dimensional structures of their complexes with the enzyme have been solved by X-ray crystallography. Differences were noted between the IC(50) values obtained for TcAChE and those for Electrophorus electricus AChE. These differences are ascribed to sequence differences in one or two residues lining the active-site gorge of the enzyme. The binding of one of the inhibitors disrupts the native conformation of one wall of the gorge, formed by the loop Trp279-Phe290. It is proposed that flexibility of this loop may permit the binding of inhibitors such as galanthamine, which are too bulky to penetrate the narrow neck of the gorge formed by Tyr121 and Phe330 as seen in the crystal structure.
- Departments of Structural Biology and Neurobiology, Weizmann Institute of Science, Rehovot 76100, Israel.
Organizational Affiliation: