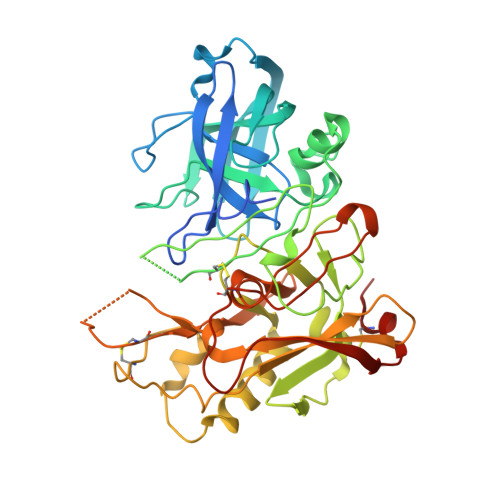
Apo and Inhibitor Complex Structures of Bace (Beta-Secretase)
Patel, S., Vuillard, L., Cleasby, A., Murray, C.W., Yon, J.(2004) J Mol Biology 343: 407
- PubMed: 15451669 Search on PubMed
- DOI: https://doi.org/10.1016/j.jmb.2004.08.018
- Primary Citation Related Structures:
1W50, 1W51 - PubMed Abstract:
Human BACE, also known as beta-secretase, shows promise as a potential therapeutic target for Alzheimer's disease. We determined the apo structure of BACE to 1.75 A, and a structure of a hydroxyethylamine inhibitor complex derived by soaking. These show significant active-site movements compared to previously described BACE structures. Additionally, the structures reveal two pockets that could be targeted by structure-based drug design.
- Astex Technology, 436 Cambridge Science Park, Milton Road, CB4 0QA, UK.
Organizational Affiliation: