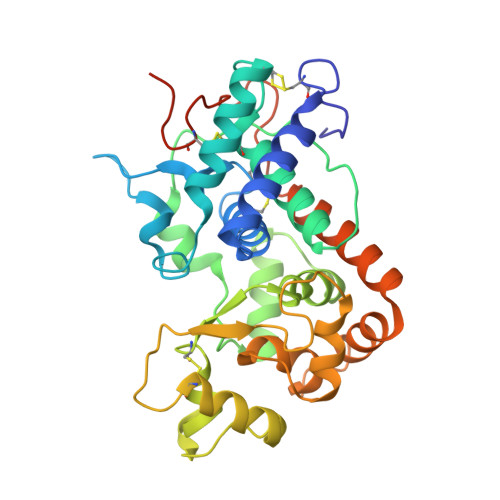
Complexes of Horseradish Peroxidase with Formate, Acetate, and Carbon Monoxide
Carlsson, G.H., Nicholls, P., Svistunenko, D., Berglund, G.I., Hajdu, J.(2005) Biochemistry 44: 635
- PubMed: 15641789 Search on PubMed
- DOI: https://doi.org/10.1021/bi0483211
- Primary Citation Related Structures:
1W4W, 1W4Y - PubMed Abstract:
Carbon monoxide, formate, and acetate interact with horseradish peroxidase (HRP) by binding to subsites within the active site. These ligands also bind to catalases, but their interactions are different in the two types of enzymes. Formate (notionally the "hydrated" form of carbon monoxide) is oxidized to carbon dioxide by compound I in catalase, while no such reaction is reported to occur in HRP, and the CO complex of ferrocatalase can only be obtained indirectly. Here we describe high-resolution crystal structures for HRP in its complexes with carbon monoxide and with formate, and compare these with the previously determined HRP-acetate structure [Berglund, G. I., et al. (2002) Nature 417, 463-468]. A multicrystal X-ray data collection strategy preserved the correct oxidation state of the iron during the experiments. Absorption spectra of the crystals and electron paramagnetic resonance data for the acetate and formate complexes in solution correlate electronic states with the structural results. Formate in ferric HRP and CO in ferrous HRP bind directly to the heme iron with iron-ligand distances of 2.3 and 1.8 A, respectively. CO does not bind to the ferric iron in the crystal. Acetate bound to ferric HRP stacks parallel with the heme plane with its carboxylate group 3.6 A from the heme iron, and without an intervening solvent molecule between the iron and acetate. The positions of the oxygen atoms in the bound ligands outline a potential access route for hydrogen peroxide to the iron. We propose that interactions in this channel ensure deprotonation of the proximal oxygen before binding to the heme iron.
- Molecular Biophysics, Institute of Cellular and Molecular Biology, Uppsala University, Husargatan 3 (Box 596), SE-75124 Uppsala, Sweden.
Organizational Affiliation: