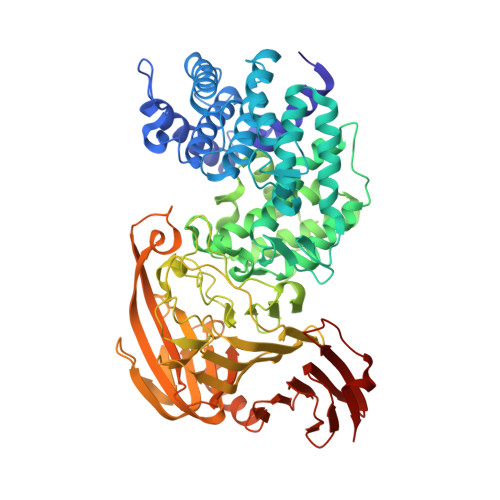
L-Ascorbic Acid 6-Hexadecanoate, a Potent Hyaluronidase Inhibitor: X-Ray Structure and Molecular Modeling of Enzyme-Inhibitor Complexes
Botzki, A., Rigden, D.J., Braun, S., Nukui, M., Salmen, S., Hoechstetter, J., Bernhardt, G., Dove, S., Jedrzejas, M.J., Buschauer, A.(2004) J Biological Chem 279: 45990
- PubMed: 15322107
- DOI: https://doi.org/10.1074/jbc.M406146200
- Primary Citation of Related Structures:
1W3Y - PubMed Abstract:
Hyaluronidases are enzymes that degrade hyaluronan, an important component of the extracellular matrix. The mammalian hyaluronidases are considered to be involved in many (patho)physiological processes like fertilization, tumor growth, and metastasis. Bacterial hyaluronidases, also termed hyaluronate lyases, contribute to the spreading of microorganisms in tissues. Such roles for hyaluronidases suggest that inhibitors could be useful pharmacological tools. Potent and selective inhibitors are not known to date, although L-ascorbic acid has been reported to be a weak inhibitor of Streptococcus pneumoniae hyaluronate lyase (SpnHL). The x-ray structure of SpnHL complexed with L-ascorbic acid has been elucidated suggesting that additional hydrophobic interactions might increase inhibitory activity. Here we show that L-ascorbic acid 6-hexadecanoate (Vcpal) is a potent inhibitor of both streptococcal and bovine testicular hyaluronidase (BTH). Vcpal showed strong inhibition of Streptococcus agalactiae hyaluronate lyase with an IC(50) of 4 microM and weaker inhibition of SpnHL and BTH with IC(50) values of 100 and 56 microM, respectively. To date, Vcpal has proved to be one of the most potent inhibitors of hyaluronidase. We also determined the x-ray structure of the SpnHL-Vcpal complex and confirmed the hypothesis that additional hydrophobic interactions with Phe-343, His-399, and Thr-400 in the active site led to increased inhibition. A homology structural model of BTH was also generated to suggest binding modes of Vcpal to this hyaluronidase. The long alkyl chain seemed to interact with an extended, hydrophobic channel formed by mostly conserved amino acids Ala-84, Leu-91, Tyr-93, Tyr-220, and Leu-344 in BTH.
- Institute of Pharmacy, University of Regensburg, Universitätsstrasse 31, 93040 Regensburg, Germany.
Organizational Affiliation: