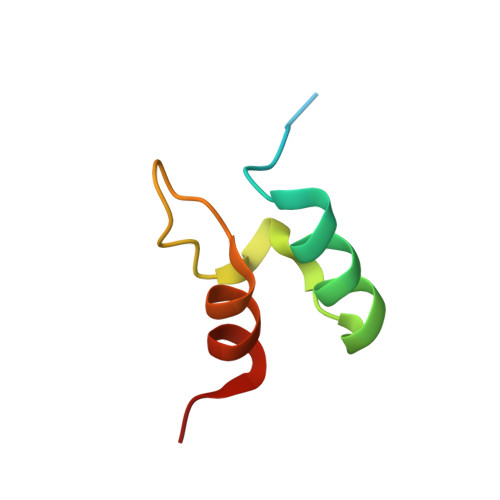
Interaction of the E2 and E3 Components of the Pyruvate Dehydrogenase Multienzyme Complex of Bacillus Stearothermophilus. Use of a Truncated Protein Domain in NMR Spectroscopy
Allen, M.D., Broadhurst, R.W., Solomon, R.G., Perham, R.N.(2005) FEBS J 272: 259
- PubMed: 15634348
- DOI: https://doi.org/10.1111/j.1432-1033.2004.04405.x
- Primary Citation of Related Structures:
1W3D - PubMed Abstract:
A (15)N-labelled peripheral-subunit binding domain (PSBD) of the dihydrolipoyl acetyltransferase (E2p) and the dimer of a solubilized interface domain (E3int) derived from the dihydrolipoyl dehydrogenase (E3) were used to investigate the basis of the interaction of E2p with E3 in the assembly of the pyruvate dehydrogenase multienzyme complex of Bacillus stearothermophilus. Thirteen of the 55 amino acids in the PSBD show significant changes in either or both of the (15)N and (1)H amide chemical shifts when the PSBD forms a 1 : 1 complex with E3int. All of the 13 amino acids reside near the N-terminus of helix I of PSBD or in the loop region between helix II and helix III. (15)N backbone dynamics experiments on PSBD indicate that the structured region extends from Val129 to Ala168, with limited structure present in residues Asn126 to Arg128. The presence of structure in the region before helix I was confirmed by a refinement of the NMR structure of uncomplexed PSBD. Comparison of the crystal structure of the PSBD bound to E3 with the solution structure of uncomplexed PSBD described here indicates that the PSBD undergoes almost no conformational change upon binding to E3. These studies exemplify and validate the novel use of a solubilized, truncated protein domain in overcoming the limitations of high molecular mass on NMR spectroscopy.
- Cambridge Centre for Molecular Recognition, Department of Biochemistry, University of Cambridge, UK.
Organizational Affiliation: