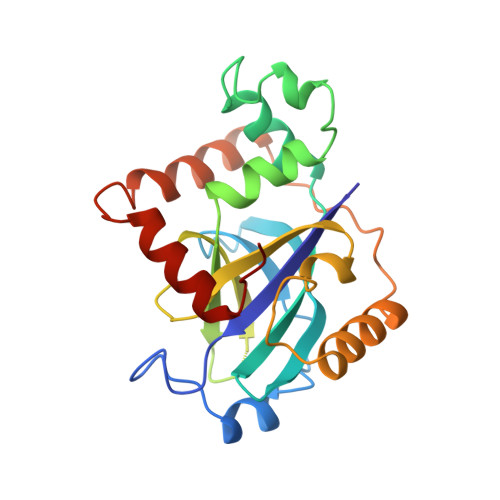
Crystal structure of varicella-zoster virus protease.
Qiu, X., Janson, C.A., Culp, J.S., Richardson, S.B., Debouck, C., Smith, W.W., Abdel-Meguid, S.S.(1997) Proc Natl Acad Sci U S A 94: 2874-2879
- PubMed: 9096314
- DOI: https://doi.org/10.1073/pnas.94.7.2874
- Primary Citation Related Structures:
1VZV - PubMed Abstract:
Varicella-zoster virus (VZV), an alpha-herpes virus, is the causative agent of chickenpox, shingles, and postherpetic neuralgia. The three-dimensional crystal structure of the serine protease from VZV has been determined at 3.0-A resolution. The VZV protease is essential for the life cycle of the virus and is a potential target for therapeutic intervention. The structure reveals an overall fold that is similar to that recently reported for the serine protease from cytomegalovirus (CMV), a herpes virus of the beta subfamily. The VZV protease structure provides further evidence to support the finding that herpes virus proteases have a fold and active site distinct from other serine proteases. The VZV protease catalytic triad consists of a serine and two histidines. The distal histidine is proposed to properly orient the proximal histidine. The identification of an alpha-helical segment in the VZV protease that was mostly disordered in the CMV protease provides a better definition of the postulated active site cavity and reveals an elastase-like S' region. Structural differences between the VZV and CMV proteases also suggest potential differences in their oligomerization states.
- Department of Macromolecular Sciences, SmithKline Beecham Pharmaceuticals, King of Prussia, PA 19406, USA.
Organizational Affiliation: