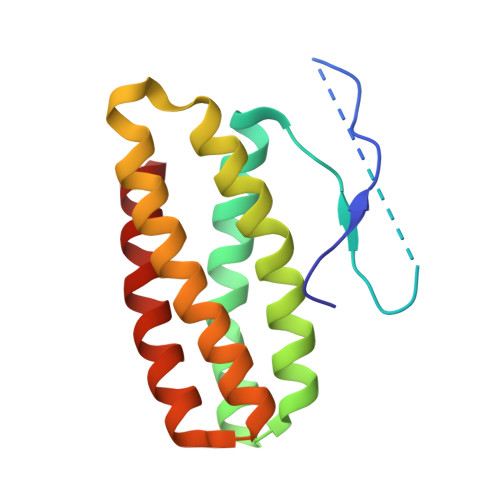
The 1.49 A Resolution Crystal Structure of Psbq from Photosystem II of Spinacia Oleracea Reveals a Ppii Structure in the N-Terminal Region.
Balsera, M., Arellano, J.B., Revuelta, J.L., De Las Rivas, J., Hermoso, J.A.(2005) J Mol Biology 350: 1051
- PubMed: 15982665
- DOI: https://doi.org/10.1016/j.jmb.2005.05.044
- Primary Citation Related Structures:
1VYK - PubMed Abstract:
We report the high-resolution structure of the spinach PsbQ protein, one of the main extrinsic proteins of higher plant photosystem II (PSII). The crystal structure shows that there are two well-defined regions in PsbQ, the C-terminal region (residues 46-149) folded as a four helix up-down bundle and the N-terminal region (residues 1-45) that is loosely packed. This structure provides, for the first time, insights into the crucial N-terminal region. First, two parallel beta-strands cross spatially, joining the beginning and the end of the N-terminal region of PsbQ. Secondly, the residues Pro9-Pro10-Pro11-Pro12 form a left-handed helix (or a polyproline type II (PPII) structure), which is stabilized by hydrogen bonds between the Pro peptide carbonyl groups and solvent water molecules. Thirdly, residues 14-33 are not visible in the electron density map, suggesting that this loop might be very flexible and presumably extended when PsbQ is free in solution. On the basis of the essential role of the N-terminal region of PsbQ in binding to PSII, we propose that both the PPII structure and the missing loop are key secondary structure elements in the recognition of specific protein-protein interactions between PsbQ and other oxygen-evolving complex extrinsic and/or intrinsic proteins of PSII. In addition, the PsbQ crystal coordinates two zinc ions, one of them is proposed to have a physiological role in higher plants, on the basis of the full conservation of the ligand protein residues in the sequence subfamily.
- Instituto de Recursos Naturales y Agrobiología (CSIC), Cordel de Merinas 52, 37008 Salamanca, Spain.
Organizational Affiliation: