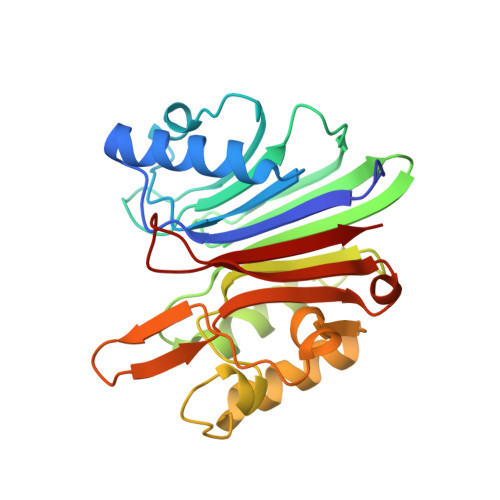
Crystal structure of the targeting endonuclease of the human LINE-1 retrotransposon.
Weichenrieder, O., Repanas, K., Perrakis, A.(2004) Structure 12: 975-986
- PubMed: 15274918 Search on PubMed
- DOI: https://doi.org/10.1016/j.str.2004.04.011
- Primary Citation Related Structures:
1VYB - PubMed Abstract:
The human L1 endonuclease (L1-EN) is encoded by the non-LTR retrotransposon LINE-1 (L1). L1 is responsible for more than 1.5 million retrotransposition events in the history of the human genome, contributing more than a quarter to human genomic DNA (L1 and Alu elements). L1-EN is related to the well-understood human DNA repair endonuclease APE1, and its nicking specificity is a major determinant for retrotransposon integration site selection. The crystal structure of human L1 endonuclease is the first of a retrotransposon-encoded protein and a prototype for retrotransposon-encoded endonucleases involved in target-primed reverse transcription. Structure-based endonuclease alignments reveal a conserved threonine in addition to previously identified invariant residues and suggest that DNA recognition proceeds via the accommodation of an extrahelical nucleotide within a pocket of the enzyme. The present analysis will help to refine phylogenetic and functional relationships among metal-dependent phosphohydrolases and provides a basis for manipulating non-LTR retrotransposon integration site selection.
- The Netherlands Cancer Institute, Department of Molecular Carcinogenesis-H2, Plesmanlaan 121, 1066 CX Amsterdam, The Netherlands. o.weichenrieder@nki.nl
Organizational Affiliation: