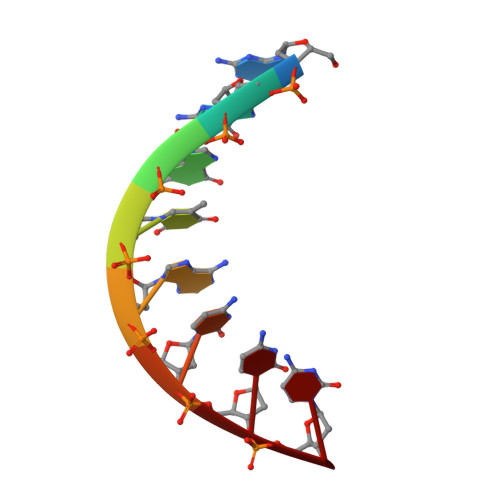
The structure and hydration of the A-DNA fragment d(GGGTACCC) at room temperature and low temperature.
Eisenstein, M., Frolow, F., Shakked, Z., Rabinovich, D.(1990) Nucleic Acids Res 18: 3185-3194
- PubMed: 2356117 Search on PubMedSearch on PubMed Central
- DOI: https://doi.org/10.1093/nar/18.11.3185
- Primary Citation Related Structures:
1VT9, 1VTA - PubMed Abstract:
The DNA fragment d(GGGTACCC) was crystallized as an A-DNA duplex in the hexagonal space group P6(1). The structure was analyzed at room temperature and low temperature (100K) at a resolution of 2.5 A. The helical conformations at the two temperatures are similar but the low-temperature structure is more economically hydrated than the room-temperature one. The structure of d(GGGTACCC) is compared to those of d(GGGTGCCC) and d(GGGCGCCC). This series of molecules, which consists of a mismatched duplex and its two Watson-Crick analogues, exhibits three conformational variants of the A-form of DNA, which are correlated with the specific intermolecular interactions observed in the various crystals. The largest differences in local conformation are displayed by the stacking geometries of the central pyrimidine-purine and the flanking purine-pyrimidine sites in each of the three duplexes. Stacking energy calculations performed on the crystal structures show that the mismatched duplex is destabilized with respect to each of the error-free duplexes, in accordance with helix-coil transition measurements.
- Department of Structural Chemistry, Weizmann Institute of Science, Rehovot, Israel.
Organizational Affiliation: