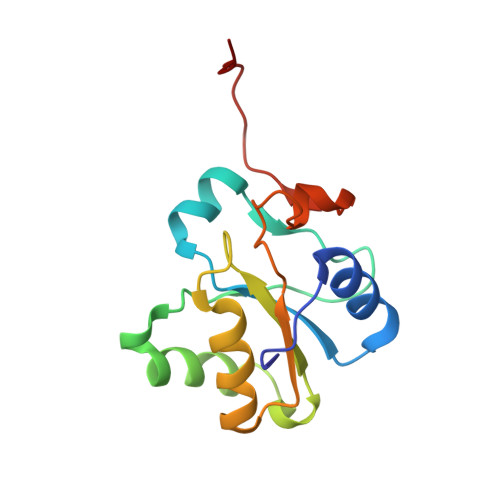
Solution structure of the rhodanese homology domain At4g01050(175-295) from Arabidopsis thaliana
Pantoja-Uceda, D., Lopez-Mendez, B., Koshiba, S., Inoue, M., Kigawa, T., Terada, T., Shirouzu, M., Tanaka, A., Seki, M., Shinozaki, K., Yokoyama, S., Guntert, P.(2005) Protein Sci 14: 224-230
- PubMed: 15576557
- DOI: https://doi.org/10.1110/ps.041138705
- Primary Citation of Related Structures:
1VEE - PubMed Abstract:
The three-dimensional structure of the rhodanese homology domain At4g01050(175-195) from Arabidopsis thaliana has been determined by solution nuclear magnetic resonance methods based on 3043 upper distance limits derived from NOE intensities measured in three-dimensional NOESY spectra. The structure shows a backbone root mean square deviation to the mean coordinates of 0.43 A for the structured residues 7-125. The fold consists of a central parallel beta-sheet with five strands in the order 1-5-4-2-3 and arranged in the conventional counterclockwise twist, and helices packing against each side of the beta-sheet. Comparison with the sequences of other proteins with a rhodanese homology domain in Arabidopsis thaliana indicated residues that could play an important role in the scaffold of the rhodanese homology domain. Finally, a three-dimensional structure comparison of the present noncatalytic rhodanese homology domain with the noncatalytic rhodanese domains of sulfurtransferases from other organisms discloses differences in the length and conformation of loops that could throw light on the role of the noncatalytic rhodanese domain in sulfurtransferases.
- Tatsuo Miyazawa Memorial Program, Japan.
Organizational Affiliation: