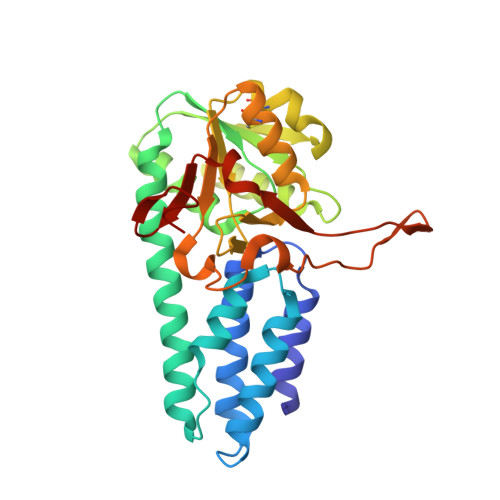
Crystal structure of the regulatory subunit of archaeal initiation factor 2B (aIF2B) from hyperthermophilic archaeon Pyrococcus horikoshii OT3: a proposed structure of the regulatory subcomplex of eukaryotic IF2B
Kakuta, Y., Tahara, M., Maetani, S., Yao, M., Tanaka, I., Kimura, M.(2004) Biochem Biophys Res Commun 319: 725-732
- PubMed: 15184043 Search on PubMed
- DOI: https://doi.org/10.1016/j.bbrc.2004.05.045
- Primary Citation Related Structures:
1VB5 - PubMed Abstract:
Eukaryotic translation initiation factor 2B (eIF2B) is the guanine-nucleotide exchange factor for eukaryotic initiation factor 2 (eIF2). eIF2B is a heteropentameric protein composed of alpha- subunits. The alpha, beta, and delta subunits form a regulatory subcomplex, while the gamma and form a catalytic subcomplex. Archaea possess homologues of alpha, beta, and delta subunits of eIF2B. Here, we report the three-dimensional structure of an archaeal regulatory subunit (aIF2Balpha) from the hyperthermophilic archaeon Pyrococcus horikoshii OT3 determined by X-ray crystallography at 2.2A resolution. aIF2Balpha consists of two subdomains, an N-domain (residues 1-95) and a C-domain (residues 96-276), connected by a long alpha-helix (alpha5: 78-106). The N-domain contains a five helix bundle structure, while the C-domain folds into the alpha/beta structure, thus showing similarity to D-ribose-5-phosphate isomerase structure. The presence of two molecules in the crystallographic asymmetric unit and the gel filtration analysis suggest a dimeric structure of aIF2Balpha in solution, interacting with each other by C-domains. Furthermore, the crystallographic 3-fold symmetry generates a homohexameric structure of aIF2Balpha; the interaction is primarily mediated by the long alpha-helix at the N-domains. This structure suggests an architecture of the three subunits, alpha, beta, and delta, in the regulatory subcomplex within eIF2B.
- Laboratory of Biochemistry, Department of Bioscience and Biotechnology, Faculty of Agriculture, Graduate School, Kyushu University, Fukuoka 812-8581, Japan.
Organizational Affiliation: