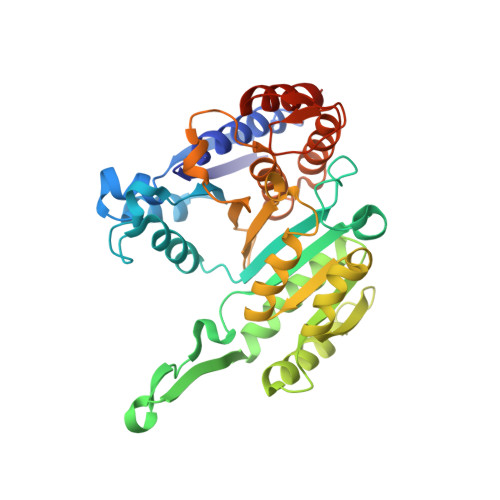
Crystal structure of a highly thermo-stabilized mutant of 3-isopropylmalate dehydrogenase from Bacillus coagulans: An evaluation of local packing density in the hydrophobic core
Fujita, K., Tsuchiya, D., Adachi, W., Suzuki, K., Tsunoda, M., Minami, H., Sekiguchi, T., Kobayashi, N., Mizui, R., Tsuzaki, S., Nakamura, S., Takenaka, A.To be published.