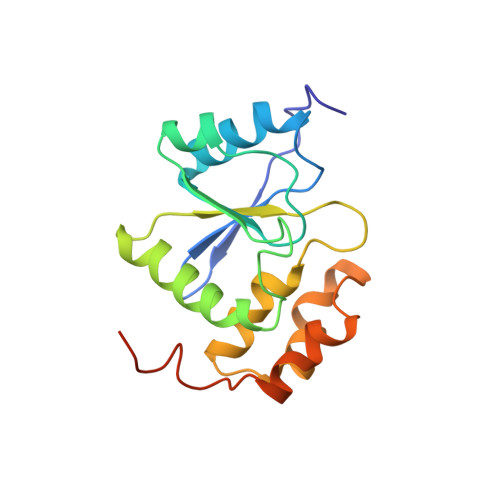
Structure of human PRL-3, the phosphatase associated with cancer metastasis
Kim, K.A., Song, J.S., Jee, J., Sheen, M.R., Lee, C., Lee, T.G., Ro, S., Cho, J.M., Lee, W., Yamazaki, T., Jeon, Y.H., Cheong, C.(2004) FEBS Lett 565: 181-187
- PubMed: 15135076
- DOI: https://doi.org/10.1016/j.febslet.2004.03.062
- Primary Citation of Related Structures:
1V3A - PubMed Abstract:
PRL-3, a novel class protein of prenylated tyrosine phosphatase, is important in cancer metastasis. Due to its high levels of expression in metastatic tumors, PRL-3 may constitute a useful marker for metastasis and might be a new therapeutic target. Here, we present the solution structure of the phosphatase domain of a human PRL-3 (residues 1-162) in phosphate-free state. The nuclear magnetic resonance (NMR) structure of PRL-3 is similar to that of other known phosphatases with minor differences in the secondary structure. But the conformation and flexibility of the loops comprising the active site differ significantly. When phosphate ions or sodium orthovanadate, which is a known inhibitor, are added to the apo PRL-3, the NMR signals from the residues in the active site appeared and could be assigned, indicating that the conformation of the residues has been stabilized.
- Magnetic Resonance Team, Korea Basic Science Institute, Daejeon 305-333, Republic of Korea.
Organizational Affiliation: