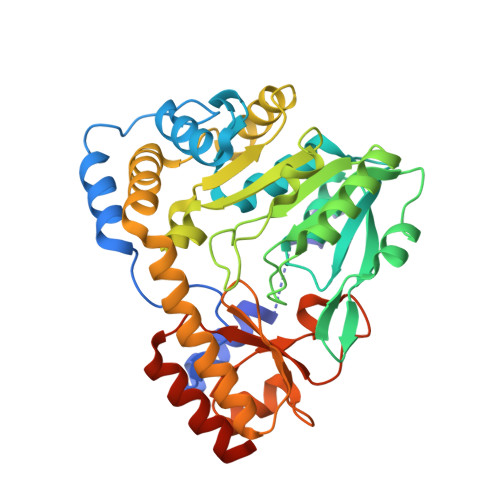
Crystal structures of glutamine:phenylpyruvate aminotransferase from Thermus thermophilus HB8: induced fit and substrate recognition
Goto, M., Omi, R., Miyahara, I., Hosono, A., Mizuguchi, H., Hayashi, H., Kagamiyama, H., Hirotsu, K.(2004) J Biological Chem 279: 16518-16525
- PubMed: 14761974
- DOI: https://doi.org/10.1074/jbc.M311575200
- Primary Citation Related Structures:
1V2D, 1V2E, 1V2F - PubMed Abstract:
The following three-dimensional structures of three forms of glutamine:phenylpyruvate aminotransferase from Thermus thermophilus HB8 have been determined and represent the first x-ray analysis of the enzyme: the unliganded pyridoxal 5'-phosphate form at 1.9 A resolution and two complexes with 3-phenylpropionate and alpha-keto-gamma-methylthiobutyrate at 2.35 and 2.6 A resolution, respectively. The enzyme shows high activity toward phenylalanine, tyrosine, tryptophan, kynurenine, methionine, and glutamine. The enzyme is a homodimer, and each subunit is divided into an N-terminal arm and small and large domains. Based on its folding, the enzyme belongs to fold type I, aminotransferase subclass Ib. The subclass I aminotransferases whose structures have so far been determined exhibit a large movement of the small domain region upon binding of a substrate. Similarly, the glutamine:phenylpyruvate aminotransferase undergoes a large movement in part of the small domain to close the active site. The active-site pocket has a shape and size suitable to enclose the side chain of an aromatic amino acid or that of methionine. The inner side of the pocket is mostly hydrophobic, but also has polar sites. The kynurenine complex generated by computer modeling fits the pocket of the enzyme and its hydrophilic groups interact with the polar sites of the pocket.
- Department of Chemistry, Graduate School of Science, Osaka City University, Osaka 558-8585, Japan.
Organizational Affiliation: