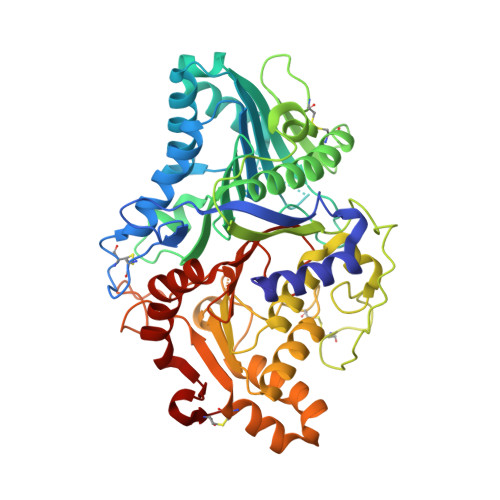
The Reaction Mechanism of Phospholipase D from Streptomyces Sp. Strain Pmf. Snapshots Along the Reaction Pathway Reveal a Pentacoordinate Reaction Intermediate and an Unexpected Final Product
Leiros, I., Mcsweeney, S., Hough, E.(2004) J Mol Biology 339: 805
- PubMed: 15165852 Search on PubMed
- DOI: https://doi.org/10.1016/j.jmb.2004.04.003
- Primary Citation Related Structures:
1V0R, 1V0S, 1V0T, 1V0U, 1V0V, 1V0W, 1V0Y - PubMed Abstract:
Almost all enzyme-catalysed phosphohydrolytic or phosphoryl transfer reactions proceed through a five-coordinated phosphorus transition state. This is also true for the phospholipase D superfamily of enzymes, where the active site usually is made up of two identical sequence repeats of an HKD motif, positioned around an approximate 2-fold axis, where the histidine and lysine residues are essential for catalysis. An almost complete reaction pathway has been elucidated by a series of experiments where crystals of phospholipase D from Streptomyces sp. strain PMF (PLD(PMF)) were soaked for different times with (i) a soluble poor, short-chained phospholipid substrate and (ii) with a product. The various crystal structures were determined to a resolution of 1.35-1.75 A for the different time-steps. Both substrate and product-structures were determined in order to identify the different reaction states and to examine if the reaction actually terminated on formation of phosphatidic acid (the true product of phospholipase D action) or could proceed even further. The results presented support the theory that the phospholipase D superfamily shares a common reaction mechanism, although different family members have very different substrate preferences and perform different catalytic reactions. Results also show that the reaction proceeds via a phosphohistidine intermediate and provide unambiguous identification of a catalytic water molecule, ideally positioned for apical attack on the phosphorus and consistent with an associative in-line phosphoryl transfer reaction. In one of the experiments an apparent five-coordinate phosphorus transition state is observed.
- Department of Chemistry, Faculty of Science, University of Tromsø, Tromsø, Norway. ingar.leiros@esrf.fr
Organizational Affiliation: