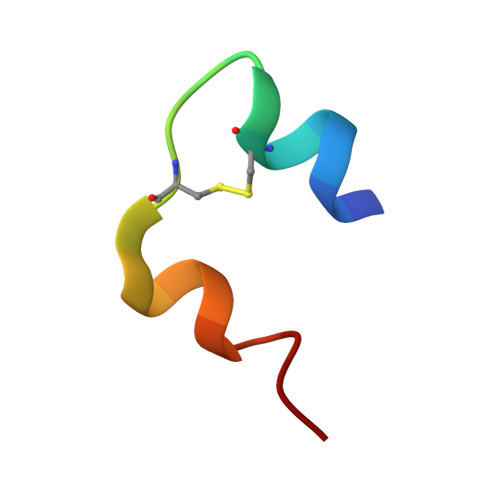
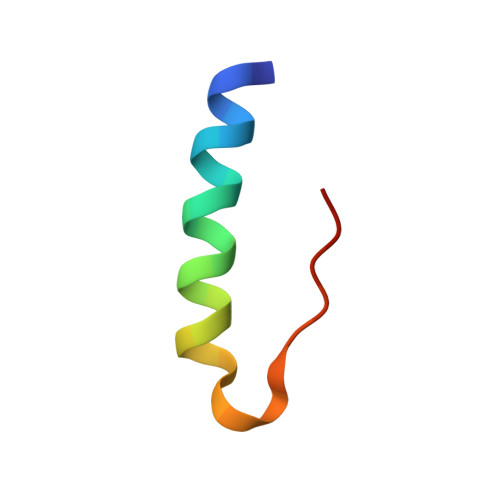
Crystallographic and Solution Studies of N-Lithocholyl Insulin: A New Generation of Prolonged-Acting Human Insulins
Whittingham, J.L., Jonassen, I., Havelund, S., Roberts, S.M., Dodson, E.J., Verma, C.S., Wilkinson, A.J., Dodson, G.G.(2004) Biochemistry 43: 5987
- PubMed: 15147182 Search on PubMed
- DOI: https://doi.org/10.1021/bi036163s
- Primary Citation Related Structures:
1UZ9 - PubMed Abstract:
The addition of specific bulky hydrophobic groups to the insulin molecule provides it with affinity for circulating serum albumin and enables it to form soluble macromolecular complexes at the site of subcutaneous injection, thereby securing slow absorption of the insulin analogue into the blood stream and prolonging its half-life once there. N-Lithocholic acid acylated insulin [Lys(B29)-lithocholyl des-(B30) human insulin] has been crystallized and the structure determined by X-ray crystallography at 1.6 A resolution to explore the molecular basis of its assembly. The unit cell in the crystal consists of an insulin hexamer containing two zinc ions, with two m-cresol molecules bound at each dimer-dimer interface stabilizing an R(6) conformation. Six covalently bound lithocholyl groups are arranged symmetrically around the outside of the hexamer. These form specific van der Waals and hydrogen-bonding interactions at the interfaces between neighboring hexamers, possibly representing the kinds of interactions which occur in the soluble aggregates at the site of injection. Comparison with an equivalent nonderivatized native insulin hexamer shows that the addition of the lithocholyl group disrupts neither the important conformational features of the insulin molecule nor its hexamer-forming ability. Indeed, binding studies show that the affinity of N-lithocholyl insulin for the human insulin receptor is not significantly diminished.
- Structural Biology Laboratory, Department of Chemistry, University of York, York YO10 5YW, UK.
Organizational Affiliation: