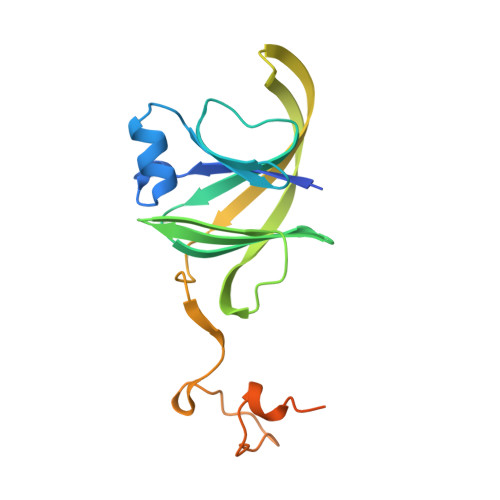
Structure and Assembly of the RNA Binding Domain of Bluetongue Virus Non-Structural Protein 2
Butan, C., Van Der Zandt, H., Tucker, P.(2004) J Biological Chem 279: 37613
- PubMed: 15155766
- DOI: https://doi.org/10.1074/jbc.M400502200
- Primary Citation of Related Structures:
1UTY - PubMed Abstract:
Bluetongue virus non-structural protein 2 belongs to a class of highly conserved proteins found in orbiviruses of the Reoviridae family. Non-structural protein 2 forms large multimeric complexes and localizes to cytoplasmic inclusions in infected cells. It is able to bind single-stranded RNA non-specifically, and it has been suggested that the protein is involved in the selection and condensation of the Bluetongue virus RNA segments prior to genome encapsidation. We have determined the x-ray structure of the N-terminal domain (sufficient for the RNA binding ability of non-structural protein 2) to 2.4 A resolution using anomalous scattering methods. Crystals of this apparently insoluble domain were obtained by in situ proteolysis of a soluble construct. The asymmetric unit shows two monomers related by non-crystallographic symmetry, with each monomer folded as a beta sandwich with a unique topology. The crystal structure reveals extensive monomer-monomer interactions, which explain the ability of the protein to self-assemble into large homomultimeric complexes. Of the entire surface area of the monomer, one-third is used to create the interfaces of the curved multimeric assembly observed in the x-ray structure. The structure reported here shows how the N-terminal domain would be able to bind single-stranded RNA non-specifically protecting the bound regions in a heterogeneous multimeric but not polymeric complex.
- European Molecular Biology Laboratory, c/o Deutsches Elektronen-Synchrotron, Notkestrasse 85, D-22603 Hamburg, Germany.
Organizational Affiliation: