Development of a Bacterial Biosensor for Nitrotoluenes: The Crystal Structure of the Transcriptional Regulator Dntr
Smirnova, I.A., Dian, C., Leonard, G.A., Mcsweeney, S., Birse, D., Brzezinski, P.(2004) J Mol Biology 340: 405
- PubMed: 15210343 Search on PubMed
- DOI: https://doi.org/10.1016/j.jmb.2004.04.071
- Primary Citation Related Structures:
1UTB, 1UTH - PubMed Abstract:
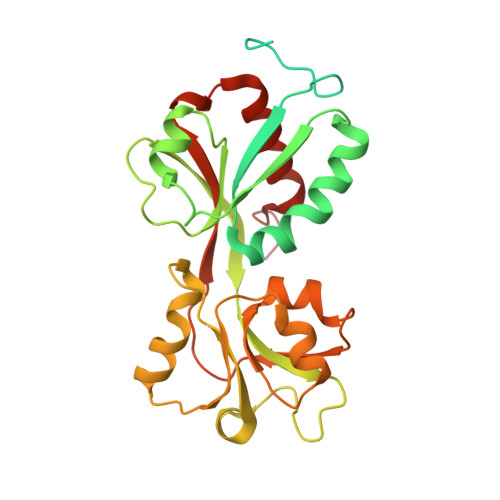
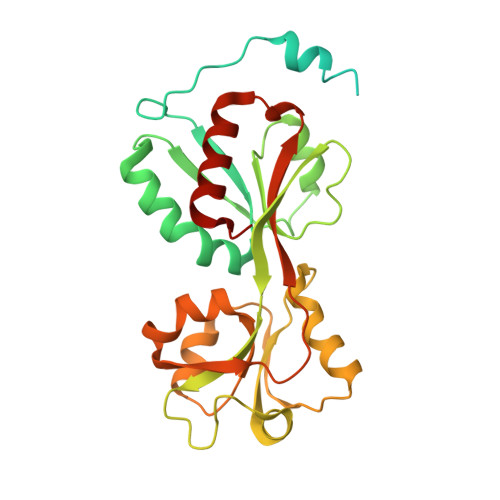
The transcriptional regulator DntR, a member of the LysR family, is a central element in a prototype bacterial cell-based biosensor for the detection of hazardous contamination of soil and groundwater by dinitrotoluenes. To optimise the sensitivity of the biosensor for such compounds we have chosen a rational design of the inducer-binding cavity based on knowledge of the three-dimensional structure of DntR. We report two crystal structures of DntR with acetate (resolution 2.6 angstroms) and thiocyanate (resolution 2.3 angstroms), respectively, occupying the inducer-binding cavity. These structures allow for the construction of models of DntR in complex with salicylate (Kd approximately or = 4 microM) and 2,4-dinitrotoluene that provide a basis for the design of mutant DntR with enhanced specificity for dinitrotoluenes. In both crystal structures DntR crystallises as a homodimer with a "head-to-tail" arrangement of monomers in the asymmetric unit. Analysis of the crystal structure has allowed the building of a full-length model of DntR in its biologically active homotetrameric form consisting of two "head-to-head" dimers. The implications of this model for the mechanism of transcription regulation by LysR proteins are discussed.
- Department of Biochemistry and Biophysics, Arrhenius Laboratories for Natural Sciences, Stockholm University, SE-106 91 Stockholm, Sweden.
Organizational Affiliation: