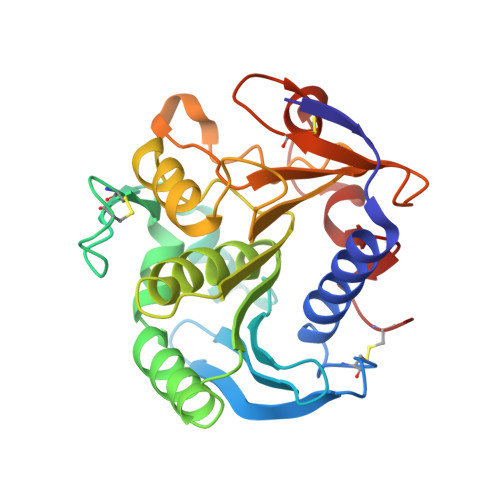
The Crystal Structure of Feruloyl Esterase a from Aspergillus Niger Suggests Evolutive Functional Convergence in Feruloyl Esterase Family
Hermoso, J., Sanz-Aparicio, J., Molina, R., Juge, N., Gonzalez, R., Faulds, C.(2004) J Mol Biology 338: 495
- PubMed: 15081808
- DOI: https://doi.org/10.1016/j.jmb.2004.03.003
- Primary Citation of Related Structures:
1USW - PubMed Abstract:
As a component of the array of enzymes produced by micro-organisms to deconstruct plant cell walls, feruloyl esterases hydrolyze phenolic groups involved in the cross-linking of arabinoxylan to other polymeric structures. This is important for opening the cell wall structure, making material more accessible to glycosyl hydrolases. Here, we describe the first crystal structure of the non-modular type-A feruloyl esterase from Aspergillus niger (AnFaeA) solved at 2.5A resolution. AnFaeA displays an alpha/beta hydrolase fold similar to that found in fungal lipases and different from that reported for other feruloyl esterases. Crystallographic and site-directed mutagenesis studies allow us to identify the catalytic triad (Ser133-His247-Asp194) that forms the catalytic machinery of this enzyme. The active-site cavity is confined by a lid (residues 68-80), on the analogy of lipases, and by a loop (residues 226-244) that confers plasticity to the substrate-binding site. The lid presents a high ratio of polar residues, which in addition to a unique N-glycosylation site stabilises the lid in an open conformation, conferring the esterase character to this enzyme. A putative model for bound 5,5'-diferulic acid-linked arabinoxylan has been built, pointing to the more relevant residues involved in substrate recognition. Comparison with structurally related lipases reveals that subtle amino acid and conformational changes within a highly conserved protein fold may produce protein variants endowed with new enzymatic properties, while comparison with functionally related proteins points to a functional convergence after evolutionary divergence within the feruloyl esterases family.
- Grupo de Cristalografía Macromolecular y Biología Estructural, Instituto Química-Física Rocasolano C.S.I.C., Serrano 119, 28006 Madrid, Spain. xjuan@iqfr.csic.es
Organizational Affiliation: