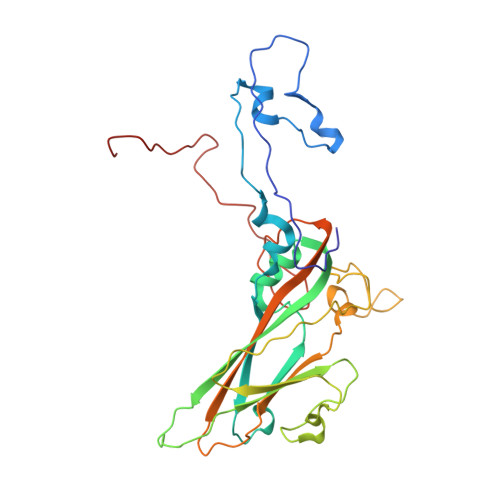
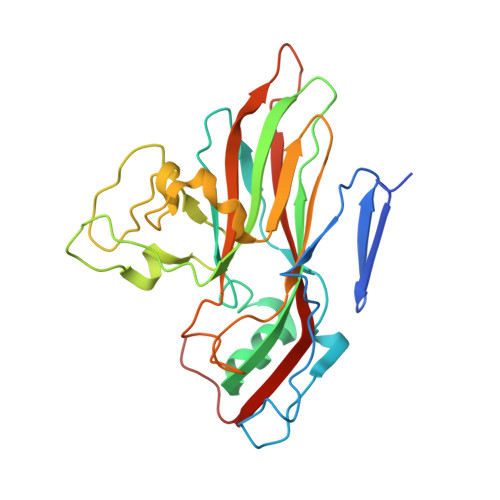
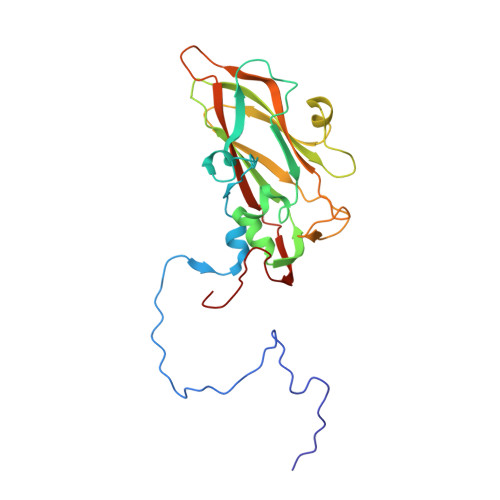
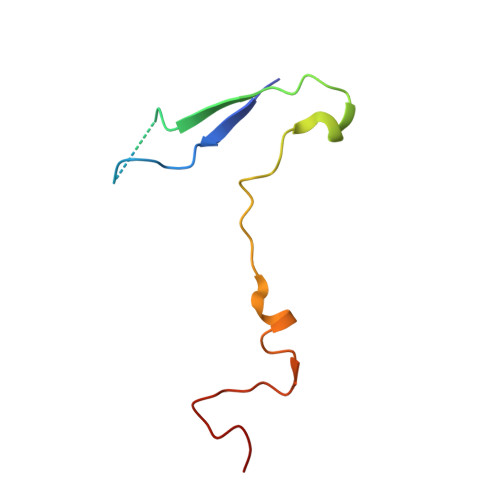
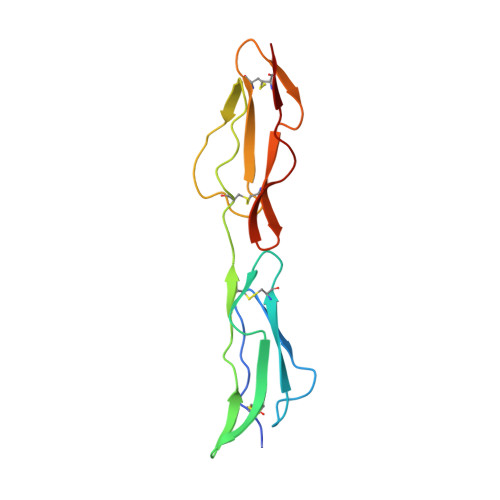
The Structure of Echovirus Type 12 Bound to a Two-Domain Fragment of its Cellular Attachment Protein Decay-Accelerating Factor (Cd 55)
Bhella, D., Goodfellow, I.G., Roversi, P., Pettigrew, D., Chaudhry, Y., Evans, D.J., Lea, S.M.(2004) J Biological Chem 279: 8325
- PubMed: 14634014 Search on PubMed
- DOI: https://doi.org/10.1074/jbc.M311334200
- Primary Citation Related Structures:
1UPN - PubMed Abstract:
Echovirus type 12 (EV12), an Enterovirus of the Picornaviridae family, uses the complement regulator decay-accelerating factor (DAF, CD55) as a cellular receptor. We have calculated a three-dimensional reconstruction of EV12 bound to a fragment of DAF consisting of short consensus repeat domains 3 and 4 from cryo-negative stain electron microscopy data (EMD code 1057). This shows that, as for an earlier reconstruction of the related echovirus type 7 bound to DAF, attachment is not within the viral canyon but occurs close to the 2-fold symmetry axes. Despite this general similarity our reconstruction reveals a receptor interaction that is quite different from that observed for EV7. Fitting of the crystallographic co-ordinates for DAF(34) and EV11 into the reconstruction shows a close agreement between the crystal structure of the receptor fragment and the density for the virus-bound receptor, allowing unambiguous positioning of the receptor with respect to the virion (PDB code 1UPN). Our finding that the mode of virus-receptor interaction in EV12 is distinct from that seen for EV7 raises interesting questions regarding the evolution and biological significance of the DAF binding phenotype in these viruses.
- Medical Research Council Virology Unit, Church Street, Glasgow, G11 5JR, United Kingdom. d.bhella@vir.gla.ac.uk
Organizational Affiliation: