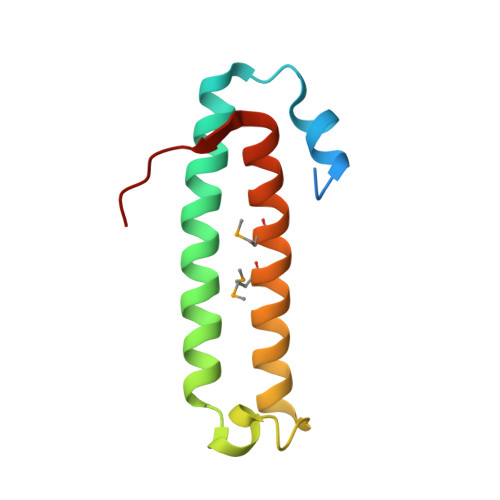
Crystal Structure of the Quorum-Sensing Protein Tram and its Interaction with the Transcriptional Regulator Trar
Vannini, A., Volpari, C., Di Marco, S.(2004) J Biological Chem 279: 24291
- PubMed: 15044488
- DOI: https://doi.org/10.1074/jbc.M401855200
- Primary Citation Related Structures:
1UPG, 1US6 - PubMed Abstract:
Transfer of the tumor-inducing plasmid in Agrobacterium tumefaciens is controlled by a quorum-sensing system whose main components are the transcriptional regulator TraR and its autoinducer. This system allows bacteria to synchronize infection of the host plant when a "quorum" of cells has been reached. TraM is an A. tumefaciens protein involved in the regulation of this system because it binds to TraR and prevents it from binding DNA. As a first step to understanding the molecular basis for the regulation of TraR by TraM, we have determined the crystal structure of TraM at 1.65 A resolution. This protein is packed as a dimer, with each monomer consisting mainly of two antiparallel alpha helices. Monomers are tightly associated, with a large hydrophobic area buried upon dimerization. Secondly, we characterized the TraR-TraM complex in vitro. TraM (11.4 kDa, monomer molecular mass) binds tightly TraR (27 kDa, monomer molecular mass) forming a stable oligomeric complex that likely accounts for two TraR and two TraM dimers.
- Department of Biochemistry, Istituto di Ricerche di Biologia Molecolare Pietro Angeletti, 00040 Pomezia, Rome, Italy.
Organizational Affiliation: