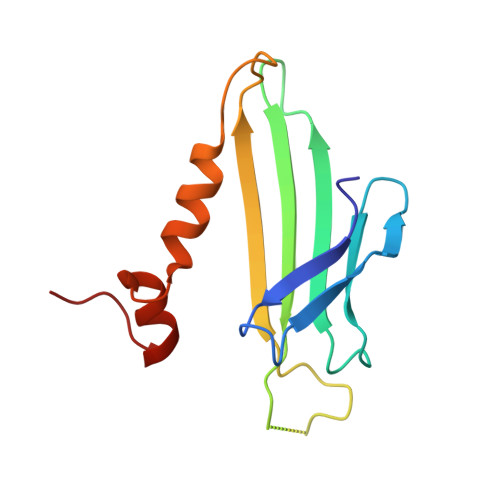
Crystal structure of the coat protein from the GA bacteriophage: model of the unassembled dimer.
Ni, C.Z., White, C.A., Mitchell, R.S., Wickersham, J., Kodandapani, R., Peabody, D.S., Ely, K.R.(1996) Protein Sci 5: 2485-2493
- PubMed: 8976557 Search on PubMedSearch on PubMed Central
- DOI: https://doi.org/10.1002/pro.5560051211
- Primary Citation Related Structures:
1UNA - PubMed Abstract:
There are four groups of RNA bacteriophages with distinct antigenic and physicochemical properties due to differences in surface residues of the viral coat proteins. Coat proteins also play a role as translational repressor during the viral life cycle, binding an RNA hairpin within the genome. In this study, the first crystal structure of the coat protein from a Group II phage GA is reported and compared to the Group I MS2 coat protein. The structure of the GA dimer was determined at 2.8 A resolution (R-factor = 0.20). The overall folding pattern of the coat protein is similar to the Group I MS2 coat protein in the intact virus (Golmohammadi R, Valegård K, Fridborg K, Liljas L. 1993, J Mol Biol 234:620-639) or as an unassembled dimer (Ni Cz, Syed R, Kodandapani R. Wickersham J, Peabody DS, Ely KR, 1995, Structure 3:255-263). The structures differ in the FG loops and in the first turn of the alpha A helix. GA and MS2 coat proteins differ in sequence at 49 of 129 amino acid residues. Sequence differences that contribute to distinct immunological and physical properties of the proteins are found at the surface of the intact virus in the AB and FG loops. There are six differences in potential RNA contact residues within the RNA-binding site located in an antiparallel beta-sheet across the dimer interface. Three differences involve residues in the center of this concave site: Lys/Arg 83, Ser/Asn 87, and Asp/Glu 89. Residue 87 was shown by molecular genetics to define RNA-binding specificity by GA or MS2 coat protein (Lim F. Spingola M, Peabody DS, 1994, J Biol Chem 269:9006-9010). This sequence difference reflects recognition of the nucleotide at position -5 in the unpaired loop of the translational operators bound by these coat proteins. In GA, the nucleotide at this position is a purine whereas in MS2, it is a pyrimidine.
- Structural Biology Program, La Jolla Cancer Research Center, Burnham Institute, California 92037, USA.
Organizational Affiliation: