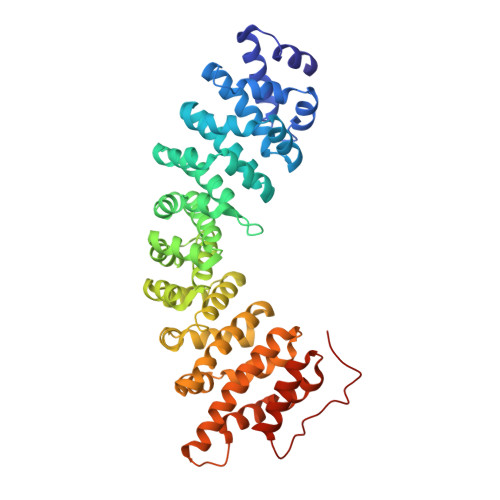
Structural Basis for Nup2P Function in Cargo Release and Karyopherin Recycling in Nuclear Import
Matsuura, Y., Lange, A., Harreman, M.T., Corbett, A.H., Stewart, M.(2003) EMBO J 22: 5358
- PubMed: 14532109 Search on PubMedSearch on PubMed Central
- DOI: https://doi.org/10.1093/emboj/cdg538
- Primary Citation Related Structures:
1UN0 - PubMed Abstract:
The yeast nucleoporin Nup2p is associated primarily with the nuclear basket of nuclear pore complexes and is required for efficient importin-alpha:beta-mediated nuclear protein import as well as efficient nuclear export of Kap60p/importin-alpha. Residues 1-51 of Nup2p bind tightly to Kap60p and are required for Nup2p function in vivo. We have determined the 2.6 A resolution crystal structure of a complex between this region of Nup2p and the armadillo repeat domain of Kap60p. Nup2p binds along the inner concave groove of Kap60p, but its interaction interface is different from that employed for nuclear localization signal (NLS) recognition although there is some overlap between them. Nup2p binds Kap60p more strongly than NLSs and accelerates release of NLSs from Kap60p. Nup2p itself is released from Kap60p by Cse1p:RanGTP only in the presence of the importin-beta binding (IBB) domain of Kap60p. These data indicate that Nup2p increases the overall rate of nuclear trafficking by coordinating nuclear import termination and importin recycling as a concerted process.
- MRC Laboratory of Molecular Biology, Hills Road, Cambridge CB2 2QH, UK.
Organizational Affiliation: