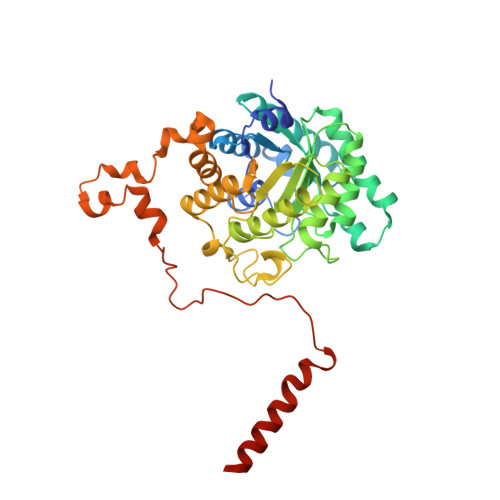
Crystal structure of rat liver betaine homocysteine s-methyltransferase reveals new oligomerization features and conformational changes upon substrate binding.
Gonzalez, B., Pajares, M.A., Martinez-Ripoll, M., Blundell, T.L., Sanz-Aparicio, J.(2004) J Mol Biology 338: 771-782
- PubMed: 15099744 Search on PubMed
- DOI: https://doi.org/10.1016/j.jmb.2004.03.005
- Primary Citation Related Structures:
1UMY - PubMed Abstract:
Betaine homocysteine S-methyltransferase (BHMT) is one of the two enzymes known to methylate homocysteine to generate methionine in the liver. It presents a Zn(2+) atom linked to three essential Cys residues. The crystal structure of rat liver BHMT has been solved at 2.5A resolution, using crystals with P2(1) symmetry and 45% solvent content in the cell. The asymmetric unit contains the whole functional tetramer showing point symmetry 222. The overall fold of the subunit consists mostly of a (alpha/beta)(8) barrel, as for human BHMT. From the end of the barrel, the polypeptide chain extends away and makes many interactions with a different subunit, forming tight dimers. The most remarkable structural feature of rat liver BHMT is the presence of a helix including residues 381-407, at the C terminus of the chain, which bind together the dimers AB to CD. A strong ion-pair and more than 60 hydrophobic interactions keep this helix stacked to the segment 316-349 from the opposite subunit. Moreover, the crystal structure of free rat liver BHMT clearly shows that Tyr160 is the fourth ligand coordinated to Zn, which is replaced by Hcy upon binding. Two residues essential for substrate recognition, Phe76 and Tyr77, are provided by a conformational change in a partially disordered loop (L2). The crucial role of these residues is highlighted by site-directed mutagenesis.
- Grupo de Cristalografía Macromolecular y Biología Estructural, Instituto de Química-Física "Rocasolano", CSIC, Serrano 119, 28006 Madrid, Spain.
Organizational Affiliation: