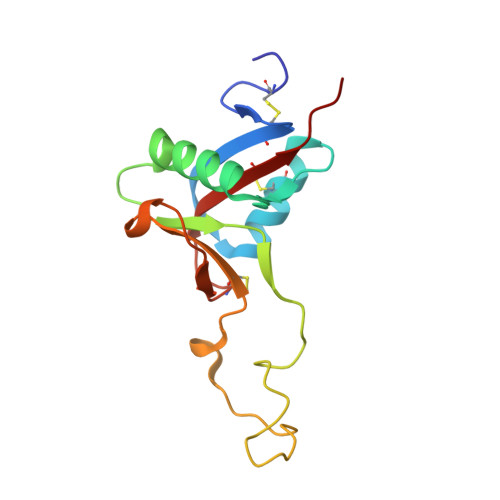
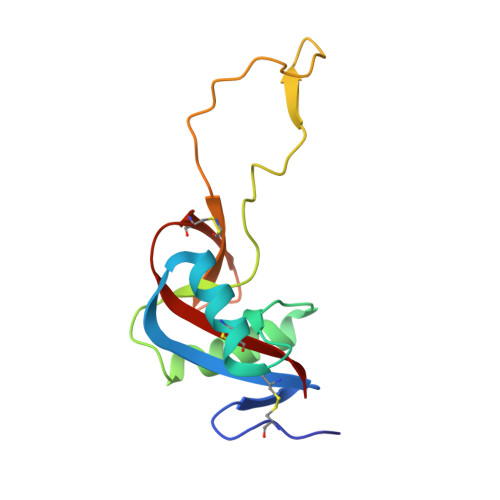
Crystal Structure of the Platelet Activator Convulxin, a Disulfide Linked A4B4 Cyclic Tetramer from the Venom of Crotalus Durissus Terrificus
Murakami, M.T., Zela, S.P., Gava, L.M., Michelan-Duarte, S., Cintra, A.C.O., Arni, R.K.(2003) Biochem Biophys Res Commun 310: 478
- PubMed: 14521935 Search on PubMed
- DOI: https://doi.org/10.1016/j.bbrc.2003.09.032
- Primary Citation Related Structures:
1UMR - PubMed Abstract:
Convulxin (CVX), a C-type lectin, isolated from the venom of the South American rattlesnake Crotalus durissus terrificus, causes cardiovascular and respiratory disturbances and is a potent platelet activator which binds to platelet glycoprotein GPVI. The structure of CVX has been solved at 2.4A resolution to a crystallographic residual of 18.6% (R(free)=26.4%). CVX is a disulfide linked heterodimer consisting of homologous alpha and beta chains. The heterodimers are additionally linked by disulfide bridges to form cyclic alpha(4)beta(4)heterotetramers. These domains exhibit significant homology to the carbohydrate-binding domains of C-type lectins, to the factor IX-binding protein (IX-bp), and to flavocetin-A (Fl-A) but sequence and structural differences are observed in both the domains in the putative Ca(2+)and carbohydrate binding regions.
- Department of Physics, IBILCE/UNESP, R. Cristovão Colombo 2265, CEP 15054-000, São Josédo Rio Preto-SP, Brazil.
Organizational Affiliation: