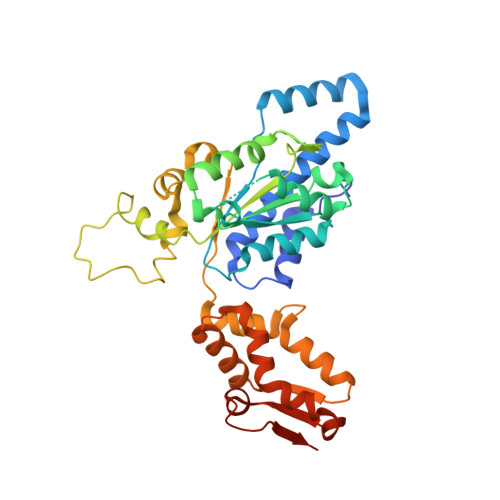
Crystal Structure of ClpX Molecular Chaperone from Helicobacter pylori
Kim, D.Y., Kim, K.K.(2003) J Biological Chem 278: 50664-50670
- PubMed: 14514695 Search on PubMed
- DOI: https://doi.org/10.1074/jbc.M305882200
- Primary Citation Related Structures:
1UM8 - PubMed Abstract:
ClpX, a heat shock protein 100 chaperone, which acts as the regulatory subunit of the ATP-dependent ClpXP protease, is responsible for intracellular protein remodeling and degradation. To provide a structural basis for a better understanding of the function of the Clp ATPase family, the crystal structures of Helicobacter pylori ClpX, lacking an N-terminal Cys cluster region complexed with ADP, was determined. The overall structure of ClpX is similar to that of heat shock locus U (HslU), consisting of two subdomains, with ADP bound at the subdomain interface. The crystal structure of ClpX reveals that a conserved tripeptide (LGF) is located on the tip of ClpP binding loop extending from the N-terminal subdomain. A hexameric model of ClpX suggests that six tripeptides make hydrophobic contacts with the hydrophobic clefts of the ClpP heptmer asymmetrically. In addition, the nucleotide binding environment provides the structural explanation for the hexameric assembly and the modulation of ATPase activity.
- Department of Molecular Cell Biology, Center for Molecular Medicine, SBRI, Sungkyunkwan University School of Medicine, Suwon 440-746, Korea.
Organizational Affiliation: