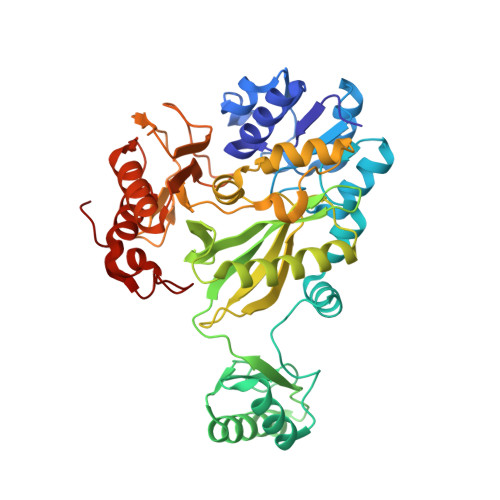
Structure of the biotin carboxylase subunit of pyruvate carboxylase from Aquifex aeolicus at 2.2 A resolution.
Kondo, S., Nakajima, Y., Sugio, S., Yong-Biao, J., Sueda, S., Kondo, H.(2004) Acta Crystallogr D Biol Crystallogr 60: 486-492
- PubMed: 14993673
- DOI: https://doi.org/10.1107/S0907444904000423
- Primary Citation Related Structures:
1ULZ - PubMed Abstract:
Pyruvate carboxylase (PC) is distributed in many eukaryotes as well as in some prokaryotes. PC catalyzes the ATP-dependent carboxylation of pyruvate to form oxalacetate. PC has three functional domains, one of which is a biotin carboxylase (BC) domain. The BC subunit of PC from Aquifex aeolicus (PC-beta) was crystallized in an orthorhombic form with space group P2(1)2(1)2, unit-cell parameters a = 92.4, b = 122.1, c = 59.0 A and one molecule in the asymmetric unit. Diffraction data were collected at 100 K on BL24XU at SPring-8. The crystal structure was determined by the molecular-replacement method and refined against 20.0-2.2 A resolution data, giving an R factor of 0.199 and a free R factor of 0.236. The crystal structure revealed that PC-beta forms a dimeric quaternary structure consisting of two molecules related by crystallographic twofold symmetry. The overall structure of PC-beta is similar to other biotin-dependent carboxylases, such as acetyl-CoA carboxylase (ACC). Although some parts of domain B were disordered in ACC, the corresponding parts of PC-beta were clearly determined in the crystal structure. From comparison between the active-site structure of ACC with ATP bound and a virtual model of PC-beta with ATP bound, it was shown that the backbone torsion angles of Glu203 in PC-beta change and some of water molecules in the active site of PC-beta are excluded upon ATP binding.
- MCC Group Science and Technology Research Center, Mitsubishi Chemical Corporation, 1000 Kamoshida-cho, Aoba-ku, Yokohama 227-8502, Japan.
Organizational Affiliation: