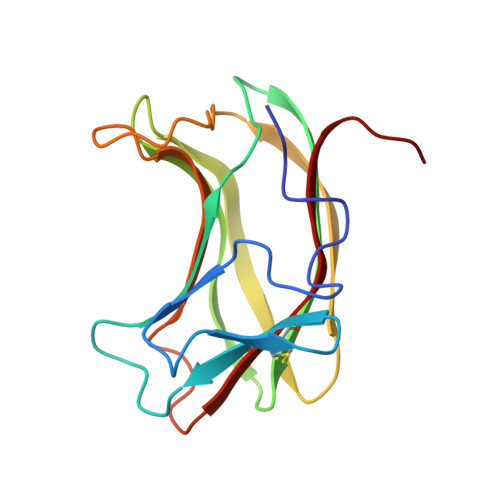
Structure of the N-terminal cellulose-binding domain of Cellulomonas fimi CenC determined by nuclear magnetic resonance spectroscopy.
Johnson, P.E., Joshi, M.D., Tomme, P., Kilburn, D.G., McIntosh, L.P.(1996) Biochemistry 35: 14381-14394
- PubMed: 8916925
- DOI: https://doi.org/10.1021/bi961612s
- Primary Citation Related Structures:
1ULO, 1ULP - PubMed Abstract:
Multidimensional heteronuclear nuclear magnetic resonance (NMR) spectroscopy was used to determine the tertiary structure of the 152 amino acid N-terminal cellulose-binding domain from Cellulomonas fimi 1,4-beta-glucanase CenC (CBDN1). CBDN1 was studied in the presence of saturating concentrations of cellotetraose, but due to spectral overlap, the oligosaccharide was not included in the structure calculations. A total of 1705 interproton nuclear Overhauser effect (NOE), 56 phi, 88 psi, 42 chi 1, 9 chi 2 dihedral angle, and 88 hydrogen-bond restraints were used to calculate 25 final structures. These structures have a rmsd from the average of 0.79 +/- 0.11 A for all backbone atoms excluding disordered termini and 0.44 +/- 0.05 A for residues with regular secondary structures. CBDN1 is composed of 10 beta-strands, folded into two antiparallel beta-sheets with the topology of a jelly-roll beta-sandwich. The strands forming the face of the protein previously determined by chemical shift perturbations to be responsible for cellooligosaccharide binding [Johnson, P. E., Tomme, P., Joshi, M. D., & McIntosh, L. P. (1996) Biochemistry 35, 13895-13906] are shorter than those forming the opposite side of the protein. This results in a 5-stranded binding cleft, containing a central strip of hydrophobic residues that is flanked on both sides by polar hydrogen-bonding groups. The presence of this cleft provides a structural explanation for the unique selectivity of CBDN1 for amorphous cellulose and other soluble oligosaccharides and the lack of binding to crystalline cellulose. The tertiary structure of CBDN1 is strikingly similar to that of the bacterial 1,3-1,4-beta-glucanases, as well as other sugar-binding proteins with jelly-roll folds.
- Protein Engineering Network of Centres of Excellence, University of British Columbia, Vancouver, Canada.
Organizational Affiliation: