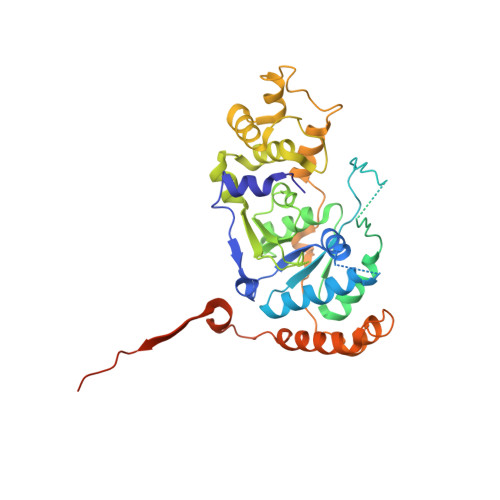
Structural basis for recruitment of human flap endonuclease 1 to PCNA
Sakurai, S., Kitano, K., Yamaguchi, H., Okada, K., Hamada, K., Fukuda, K., Uchida, M., Ohtsuka, E., Morioka, H., Hakoshima, T.(2005) EMBO J 24: 683-693
- PubMed: 15616578 Search on PubMedSearch on PubMed Central
- DOI: https://doi.org/10.1038/sj.emboj.7600519
- Primary Citation Related Structures:
1UL1 - PubMed Abstract:
Flap endonuclease-1 (FEN1) is a key enzyme for maintaining genomic stability and replication. Proliferating cell nuclear antigen (PCNA) binds FEN1 and stimulates its endonuclease activity. The structural basis of the FEN1-PCNA interaction was revealed by the crystal structure of the complex between human FEN1 and PCNA. The main interface involves the C-terminal tail of FEN1, which forms two beta-strands connected by a short helix, the betaA-alphaA-betaB motif, participating in beta-beta and hydrophobic interactions with PCNA. These interactions are similar to those previously observed for the p21CIP1/WAF1 peptide. However, this structure involving the full-length enzyme has revealed additional interfaces that are involved in the core domain. The interactions at the interfaces maintain the enzyme in an inactive 'locked-down' orientation and might be utilized in rapid DNA-tracking by preserving the central hole of PCNA for sliding along the DNA. A hinge region present between the core domain and the C-terminal tail of FEN1 would play a role in switching the FEN1 orientation from an inactive to an active orientation.
- Structural Biology Laboratory, Nara Institute of Science and Technology, Takayama, Ikoma, Nara, Japan.
Organizational Affiliation: