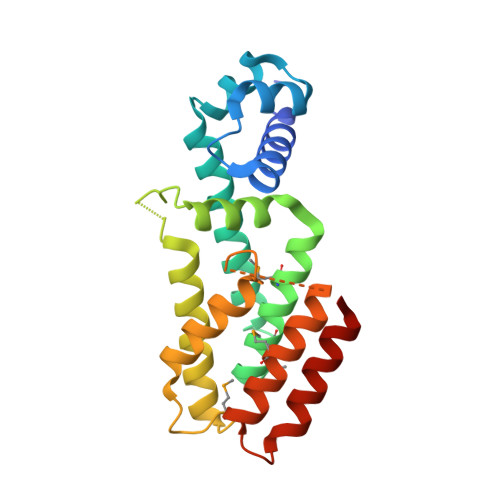
Crystal structure of a gamma-butyrolactone autoregulator receptor protein in Streptomyces coelicolor A3(2)
Natsume, R., Ohnishi, Y., Senda, T., Horinouchi, S.(2004) J Mol Biology 336: 409-419
- PubMed: 14757054
- DOI: https://doi.org/10.1016/j.jmb.2003.12.040
- Primary Citation of Related Structures:
1UI5, 1UI6 - PubMed Abstract:
The gamma-butyrolactone-type autoregulator/receptor systems in the Gram-positive bacterial genus Streptomyces regulate morphological differentiation or antibiotic production, or both. The autoregulator receptors act as DNA-binding proteins, and on binding their cognate ligands (gamma-butyrolactones) they are released from the DNA, thus serving as repressors. The crystal structure of CprB in Streptomyces coelicolor A3(2), a homologue of the A-factor-receptor protein, ArpA, in Streptomyces griseus, was determined. The overall structure of CprB shows that the gamma-butyrolactone receptors belong to the TetR family. CprB is composed of two domains, a DNA-binding domain and a regulatory domain. The regulatory domain contains a hydrophobic cavity, which probably serves as a ligand-binding pocket. On the basis of the crystal structure of CprB and on the analogy of the characteristics of ligand-TetR binding, the binding of gamma-butyrolactones to the regulatory domain of the receptors is supposed to induce the relocation of the DNA-binding domain through conformational changes of residues located between the ligand-binding site and the DNA-binding domain, which would result in the dissociation of the receptors from their target DNA.
- Department of Biotechnology, Graduate School of Agriculture and Life Sciences, The University of Tokyo, 1-1-1 Yayoi, Bunkyo-ku, 113-8657, Tokyo, Japan.
Organizational Affiliation: