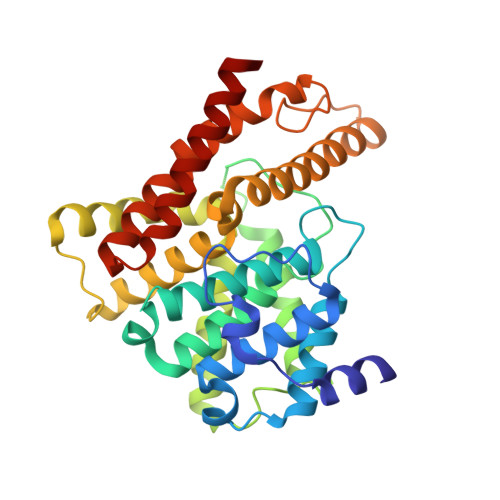
Structure of the catalytic domain of human phosphodiesterase 5 with bound drug molecules
Sung, B.-J., Hwang, K.Y., Jeon, Y.H., Lee, J.I., Heo, Y.-S., Kim, J.H., Moon, J., Yoon, J.M., Hyun, Y.-L., Kim, E., Eum, S.J., Park, S.-Y., Lee, J.-O., Lee, T.G., Ro, S., Cho, J.M.(2003) Nature 425: 98-102
- PubMed: 12955149 Search on PubMed
- DOI: https://doi.org/10.1038/nature01914
- Primary Citation Related Structures:
1UDT, 1UDU, 1UHO - PubMed Abstract:
Phosphodiesterases (PDEs) are a superfamily of enzymes that degrade the intracellular second messengers cyclic AMP and cyclic GMP. As essential regulators of cyclic nucleotide signalling with diverse physiological functions, PDEs are drug targets for the treatment of various diseases, including heart failure, depression, asthma, inflammation and erectile dysfunction. Of the 12 PDE gene families, cGMP-specific PDE5 carries out the principal cGMP-hydrolysing activity in human corpus cavernosum tissue. It is well known as the target of sildenafil citrate (Viagra) and other similar drugs for the treatment of erectile dysfunction. Despite the pressing need to develop selective PDE inhibitors as therapeutic drugs, only the cAMP-specific PDE4 structures are currently available. Here we present the three-dimensional structures of the catalytic domain (residues 537-860) of human PDE5 complexed with the three drug molecules sildenafil, tadalafil (Cialis) and vardenafil (Levitra). These structures will provide opportunities to design potent and selective PDE inhibitors with improved pharmacological profiles.
- Division of Drug Discovery, CrystalGenomics, Inc., Daedeok Biocommunity, Jeonmin-dong, Yuseong-gu, Daejeon, 305-390, South Korea.
Organizational Affiliation: